This paper came up in comments to the last post:
Dietary Stearic Acid Leads to a Reduction of Visceral Adipose Tissue in Athymic Nude Mice
I think we can say that, at least in athymic nude mice (which do not seem to be derived from the C57Bl/6 strain), omega 6 PUFA do not cause obesity when compared to either a low fat or high stearic acid synthetic diet (ie the low fat arm is equally synthetic, not more "food-like" ie not chow). At least when you look at total body weights:
So omega 6 PUFA appear to get a free pass here. The actual composition of the diets is in Table 1 of this previous paper and all four contain generous amounts of starch and equal amounts of sucrose:
Dietary Stearate Reduces Human Breast Cancer Metastasis Burden in Athymic Nude Mice
However if you dexa scan the mice you find that the low fat, corn oil and safflower oil groups all have reduced lean mass (probably muscle) and increased visceral fat mass compared to the stearate group. A picture is worth a thousand words so here are some postmortem images with the size of the inguinal fat pads outline by the authors of the paper (no need for me to doodle on this one!). Fig 3:
I really like these images.
Now, cavenewt questioned the relevance of weight/fat alterations from stearate compared to other potential health effects, particularly its affect on cancer metabolism.
The third paper from this same group is
Prevention of carcinogenesis and inhibition of breast cancer tumor burden by dietary stearate
I've been through all three papers and searched on "insulin". The group appears to have no concept that insulin has anything to do with adipocyte size or cancer progression.
A slight handicap when it comes to insight.
In the stearate-visceral fat paper there is a single measurement made of plasma insulin/glucose. Insulin does not vary between diet groups but glucose is significantly lower in the stearate group. I have been unable to work out if the measures were fasting or fed, or even what time of day the samples were taken (ie when the mice were killed). I think that with glucose values in to 200-250mg/dl range these were probably "fed" glucose and insulin levels. The paper does not give us the measured insulin levels, merely that there was no statistically significant difference between groups. But insulin levels come with such huge standard deviations that getting a p value below 0.05 with small group sizes is not going to happen. A ns result does not automatically mean that there were no differences.
Of course a single insulin measurement at one terminal time point tells us nothing about the long term 24h exposure to insulin of the mice, of their adipocytes or of their cancer cells.
So we have to, once again, look at the significance of the changes in fat distribution to attempt to gain insight in to overall insulin exposure. I spent quite some time looking at visceral fat and its significance early last year in this post:
On phosphorylation of AKT in real, live humans. They're just like mice!
and on how stearate might avoid systemic hyperinsulinaemia here:
Dairy and diabetes
Visceral fat is a surrogate for chronic hyperinsulinaemia, particularly fasting hyperinsulinaemia. While I consider non-inflamed visceral fat to be completely benign, or even beneficial for controlling the hunger of fasting, the insulin which maintains that visceral lipid storage is not benign. Chronically elevated insulin (or, more accurately, insulin signalling) should drive both visceral fat storage and xenograft tumour growth in the mice. Probably in humans too.
Happy Solstice and assorted mid-Winter celebrations. If you live in the northern hemisphere that is. Not that I envy those with a Solstice-on-the-beach-without-wooly-hats-and-gloves situation!
Peter
Tuesday, December 24, 2019
Saturday, December 14, 2019
Protons (52): A correction and a few thoughts on satiety
There has never been a completed concept of the Protons thread. Logic and data have allowed it to emerge and I still have no idea where it will end, but there have been a few mis-steps on the way. A reader recently pointed my incorrect post from 2013 where I was looking to see how fasting or ketogenic diets might blunt insulin signalling. Nowadays my feeling is that it is high physiological ROS generation which achieves this. At the time I clearly got it wrong with:
"We appear to have two basic states of the electron transport chain. There is the situation under fasting or ketogenic dieting conditions. Here delta psi is low, complex I throughput is low and there is plenty of FADH2 input through electron transporting flavoprotein dehydrogenase coming from the first step of beta oxidation of real fats, like palmitic acid. With a low delta psi it is near impossible to generate reverse electron flow through complex I so activation of insulin signalling is rapidly aborted by the continuing action of tyrosine phosphatase".
That does not appear to be the case and, having thought about it, my email reply went like this. We had been chatting about ketones, delta psi, RET and insulin resistance from ketones vs from saturated fats. I may as well copy/paste some of it here, slightly edited for clarity, and place a link within the 2013 post:
"My thinking currently is that ketones do not induce insulin resistance unless, like glucose, there is enough input via complex II to do this. So RET is possible but will only occur when the cell has adequate caloric supply. If you combined ketones with stearic or palmitic acid the long chain fatty acid would undoubtedly input at ETFdh and drive RET, rather than it being the ketones. Incidentally, this probably also represents "cellular satiety". Clearly under physiological conditions ketones will normally be associated with elevated FFAs.
The differential effects of FFAs vs ketones would be that FFAs would then drive insulin resistance in cells which can metabolise them, muscle etc, but ketones would drive far less RET in cells which require some small amount of glucose in excess of the energy supplied by the ketones themselves, ie neurones…
On the adipocytes things look more complex. Undoubtedly both ketones and FFAs exert a negative feedback on lipolysis in addition to any effect of their driving/not driving RET during oxidation. But some degree of insulin signalling is essential to physiology in addition to this negative feedback, otherwise we get diabetic ketoacidosis due to failure of insulin availability to oppose glucagon.
Once you start to think about stearic acid plus glucose you have to differentiate between cellular levels and dietary levels. Simply treating isolated adipocytes with elevated glucose and elevated long chain FFAs will result in ROS. If you set your experiment up correctly (no MUFA, no PUFA, high glucose) the resulting RET will result in apoptosis or necrosis. There is an infinite supply of papers doing this to almost any cell type.
In the whole animal things are different. If physiology is functional the stearic acid will provoke a prompt first phase insulin response and will effectively augment insulin delivery via the portal vein to the liver. Insulin acting in the liver will limit glucose release to the systemic circulation and so limit the need for systemic hyperinsulinaemia to deal with the glucose. Then the animal stays slim as there is minimal systemic hyperinsulinaemia.
If physiology is non functional we are talking DM T2 which is largely the aftermath of chronic PUFA ingestion and things then become even more complex. The low carb approach side steps the problems of failure of correct insulin secretion/hepatic signalling by simply reducing the reliance on any sort of insulin signalling beyond the most minimal needed. Then stearic acid becomes problem free. When (or if) physiology normalises then some degree of glucose consumption by mouth might be acceptable along side saturated fats but by then anyone with any sense will be fat adapted and unwilling to go back to a mixed diet".
This set of thoughts is currently relevant as Brad Marshall has taken the concept of stearic acid as the best physiological generator of adipocyte physiological insulin resistance and converted it in to a moderate carb, highly saturated fat diet with interesting results. You can read about his croissant diet on his blog Fire in a Bottle: Introducing The Croissant Diet. This experiment is based on the PhD of Valerie Reeves where a mixed macro diet based around stearic acid markedly limited weight gain in db/db mice. The db/db mouse is an extremely severe obesity model as it lacks functional leptin receptors. So a complete lack of leptin signalling can be side-stepped, still in the presence of starch, by supplying roughly 40% of calories as predominantly stearic acid. Not low carb. Not ketogenic. It works (from my point of view) by directly manipulating ROS generation (ie increasing it) at the electron transport chain level to signal "cellular" satiety, with appropriate ability to resist insulin's fat storage signal. This will be recapitulated in the brainstem neural circuits which control whole body satiety. Signalling from the ETC is core. It works by generating physiological insulin resistance and clearly over-rides any effect from leptin signalling.
Also in the comments of the last post came this gem from ctviggen, worth a post in its own right. Interesting paper. Limited insights within the discussion but great data!
Dietary Stearic Acid Leads to a Reduction of Visceral Adipose Tissue in Athymic Nude Mice
While all of these ideas were kicking around the concept of fasting as a state of caloric excess emerged. Obviously, not eating makes you hungry. Initially. There comes a point where, when insulin is low enough and FFAs rise to levels approaching over 2000micromol/l (plus ketones), when hunger decreases. Prolonged fasting does not invariably generate ravenous hunger. This is because FFAs (and ketones) represent an energy glut beyond any single cell's imagination and it does not require insulin signalling to access it (there do appear to be other controls on ATP generation). So cells which can metabolise FFAs should behave as it they have more than enough calories so that they should resist insulin. Access to excess metabolic substrate must result in ROS and the appropriate disabling of insulin signalling. Starvation as generator of a caloric excess signal... An interesting concept.
And PUFA, failing to generate adequate ROS, might well lead to glucose "wastage" in to cells such as myocytes which might well result in profound and symptomatic hypoglycaemia, essentially a failure of satiety signalling. Another interesting idea. Which clearly would not happen if metabolism was based around stearic acid...
Peter
"We appear to have two basic states of the electron transport chain. There is the situation under fasting or ketogenic dieting conditions. Here delta psi is low, complex I throughput is low and there is plenty of FADH2 input through electron transporting flavoprotein dehydrogenase coming from the first step of beta oxidation of real fats, like palmitic acid. With a low delta psi it is near impossible to generate reverse electron flow through complex I so activation of insulin signalling is rapidly aborted by the continuing action of tyrosine phosphatase".
That does not appear to be the case and, having thought about it, my email reply went like this. We had been chatting about ketones, delta psi, RET and insulin resistance from ketones vs from saturated fats. I may as well copy/paste some of it here, slightly edited for clarity, and place a link within the 2013 post:
"My thinking currently is that ketones do not induce insulin resistance unless, like glucose, there is enough input via complex II to do this. So RET is possible but will only occur when the cell has adequate caloric supply. If you combined ketones with stearic or palmitic acid the long chain fatty acid would undoubtedly input at ETFdh and drive RET, rather than it being the ketones. Incidentally, this probably also represents "cellular satiety". Clearly under physiological conditions ketones will normally be associated with elevated FFAs.
The differential effects of FFAs vs ketones would be that FFAs would then drive insulin resistance in cells which can metabolise them, muscle etc, but ketones would drive far less RET in cells which require some small amount of glucose in excess of the energy supplied by the ketones themselves, ie neurones…
On the adipocytes things look more complex. Undoubtedly both ketones and FFAs exert a negative feedback on lipolysis in addition to any effect of their driving/not driving RET during oxidation. But some degree of insulin signalling is essential to physiology in addition to this negative feedback, otherwise we get diabetic ketoacidosis due to failure of insulin availability to oppose glucagon.
Once you start to think about stearic acid plus glucose you have to differentiate between cellular levels and dietary levels. Simply treating isolated adipocytes with elevated glucose and elevated long chain FFAs will result in ROS. If you set your experiment up correctly (no MUFA, no PUFA, high glucose) the resulting RET will result in apoptosis or necrosis. There is an infinite supply of papers doing this to almost any cell type.
In the whole animal things are different. If physiology is functional the stearic acid will provoke a prompt first phase insulin response and will effectively augment insulin delivery via the portal vein to the liver. Insulin acting in the liver will limit glucose release to the systemic circulation and so limit the need for systemic hyperinsulinaemia to deal with the glucose. Then the animal stays slim as there is minimal systemic hyperinsulinaemia.
If physiology is non functional we are talking DM T2 which is largely the aftermath of chronic PUFA ingestion and things then become even more complex. The low carb approach side steps the problems of failure of correct insulin secretion/hepatic signalling by simply reducing the reliance on any sort of insulin signalling beyond the most minimal needed. Then stearic acid becomes problem free. When (or if) physiology normalises then some degree of glucose consumption by mouth might be acceptable along side saturated fats but by then anyone with any sense will be fat adapted and unwilling to go back to a mixed diet".
This set of thoughts is currently relevant as Brad Marshall has taken the concept of stearic acid as the best physiological generator of adipocyte physiological insulin resistance and converted it in to a moderate carb, highly saturated fat diet with interesting results. You can read about his croissant diet on his blog Fire in a Bottle: Introducing The Croissant Diet. This experiment is based on the PhD of Valerie Reeves where a mixed macro diet based around stearic acid markedly limited weight gain in db/db mice. The db/db mouse is an extremely severe obesity model as it lacks functional leptin receptors. So a complete lack of leptin signalling can be side-stepped, still in the presence of starch, by supplying roughly 40% of calories as predominantly stearic acid. Not low carb. Not ketogenic. It works (from my point of view) by directly manipulating ROS generation (ie increasing it) at the electron transport chain level to signal "cellular" satiety, with appropriate ability to resist insulin's fat storage signal. This will be recapitulated in the brainstem neural circuits which control whole body satiety. Signalling from the ETC is core. It works by generating physiological insulin resistance and clearly over-rides any effect from leptin signalling.
Also in the comments of the last post came this gem from ctviggen, worth a post in its own right. Interesting paper. Limited insights within the discussion but great data!
Dietary Stearic Acid Leads to a Reduction of Visceral Adipose Tissue in Athymic Nude Mice
While all of these ideas were kicking around the concept of fasting as a state of caloric excess emerged. Obviously, not eating makes you hungry. Initially. There comes a point where, when insulin is low enough and FFAs rise to levels approaching over 2000micromol/l (plus ketones), when hunger decreases. Prolonged fasting does not invariably generate ravenous hunger. This is because FFAs (and ketones) represent an energy glut beyond any single cell's imagination and it does not require insulin signalling to access it (there do appear to be other controls on ATP generation). So cells which can metabolise FFAs should behave as it they have more than enough calories so that they should resist insulin. Access to excess metabolic substrate must result in ROS and the appropriate disabling of insulin signalling. Starvation as generator of a caloric excess signal... An interesting concept.
And PUFA, failing to generate adequate ROS, might well lead to glucose "wastage" in to cells such as myocytes which might well result in profound and symptomatic hypoglycaemia, essentially a failure of satiety signalling. Another interesting idea. Which clearly would not happen if metabolism was based around stearic acid...
Peter
Wednesday, November 27, 2019
Of mice and men
I usually start from the premise that insulin makes you fat. The most simplistic prediction from this is that eating carbohydrate raises insulin and this insulin is what makes you fat. Over the years I have looked at data from all sorts of places, particularly the extremes such as the Kempner Rice Diet and/or the Potato Diet, which clearly work and which appear to do so (to me!) via lowering the level of systemic insulin acting on adipocytes. In particular I consider that first pass hepatic insulin extraction has a huge effect on the systemic insulin level and the subsequent exposure of adipocytes to that insulin. The carbohydrate-insulin-model of obesity as set up by detractors is simplistic in the extreme.
This paper was published recently:
The carbohydrate-insulin model does not explain the impact of varying dietary macronutrients on body weight and adiposity of mice
Of all of the combinations tested the one which we are interested in is where protein was held constant at 10% of calories and carbohydrate was varied from 10% of calories up to 80% of calories. The remaining calories were made up of fat giving a pure carb vs fat comparison. At the end of the study period we have this graph where I have added in the percentages of calories from fat in red along the x axis:
If you draw a straight line through the data points your weight line slopes downwards as fat percentage drops, pretty well. Fat makes you fat:
Now, clearly, there is a missing data point. That is the bodyweight from a diet group with zero carbohydrate, 10% protein and 90% fat. This combination was not included in the study.
So all else from here forward in this post is now pure speculation.
Help is at hand for the missing data point in this paper
A high-fat, ketogenic diet induces a unique metabolic state in mice
Here we also have C57Bl/6 mice and in this case they were fed F3666 diet which looks like this:
"The proportion of calories deriving from different nutrients was as follows: ... KD:95% fat, 0% carbohydrate (0% sucrose), 5% protein"
This is not a perfect fit as the protein is about half that used by Speakman and there is (obviously) no sucrose, but it's the best anyone can do in the absence of the omitted group essential to complete the graph. The mice which were eventually put on to F3666 were initially made obese with a sucrose/fat combination before being put on to their ketogenic diet. Their weight dropped from approx 37g on the sucrose/fat diet to approx 27g on F3666, ie they ended up about 3g lower in total bodyweight than the control mice fed approx 10% of calories from fat in a carbohydrate based diet throughout. Here's the graph we all know from years ago. Open triangles show the drop in weight when F3666 was introduced at around day 80:
So what might a zero carb, fairly low protein group of C57Bl/6 mice look like? They might well end up slightly below the weight of Speakman's 80% carb group mice, (ie those eating closest to mouse chow), or they might end up slightly heavier due to the higher protein content limiting the total fat percentage which could be provided. I feel a compromise might be to use 35g, the same as Speakman's 80% carb group, the closest chow equivalent.
I've added the zero carb speculative data point to the blue line on the graph at 35g bodyweight which now looks like this:
and now we can curve fit the bodyweights like this:
Ah, that's better. Ketosis at the left, "carbosis" at the right. Nice.
I love rodent studies. You just have to understand that setting them up correctly is essential to obtaining the result you want. You also have to know what you want.
Peter
This paper was published recently:
The carbohydrate-insulin model does not explain the impact of varying dietary macronutrients on body weight and adiposity of mice
Of all of the combinations tested the one which we are interested in is where protein was held constant at 10% of calories and carbohydrate was varied from 10% of calories up to 80% of calories. The remaining calories were made up of fat giving a pure carb vs fat comparison. At the end of the study period we have this graph where I have added in the percentages of calories from fat in red along the x axis:
If you draw a straight line through the data points your weight line slopes downwards as fat percentage drops, pretty well. Fat makes you fat:
Now, clearly, there is a missing data point. That is the bodyweight from a diet group with zero carbohydrate, 10% protein and 90% fat. This combination was not included in the study.
So all else from here forward in this post is now pure speculation.
Help is at hand for the missing data point in this paper
A high-fat, ketogenic diet induces a unique metabolic state in mice
Here we also have C57Bl/6 mice and in this case they were fed F3666 diet which looks like this:
"The proportion of calories deriving from different nutrients was as follows: ... KD:95% fat, 0% carbohydrate (0% sucrose), 5% protein"
This is not a perfect fit as the protein is about half that used by Speakman and there is (obviously) no sucrose, but it's the best anyone can do in the absence of the omitted group essential to complete the graph. The mice which were eventually put on to F3666 were initially made obese with a sucrose/fat combination before being put on to their ketogenic diet. Their weight dropped from approx 37g on the sucrose/fat diet to approx 27g on F3666, ie they ended up about 3g lower in total bodyweight than the control mice fed approx 10% of calories from fat in a carbohydrate based diet throughout. Here's the graph we all know from years ago. Open triangles show the drop in weight when F3666 was introduced at around day 80:
So what might a zero carb, fairly low protein group of C57Bl/6 mice look like? They might well end up slightly below the weight of Speakman's 80% carb group mice, (ie those eating closest to mouse chow), or they might end up slightly heavier due to the higher protein content limiting the total fat percentage which could be provided. I feel a compromise might be to use 35g, the same as Speakman's 80% carb group, the closest chow equivalent.
I've added the zero carb speculative data point to the blue line on the graph at 35g bodyweight which now looks like this:
and now we can curve fit the bodyweights like this:
Ah, that's better. Ketosis at the left, "carbosis" at the right. Nice.
I love rodent studies. You just have to understand that setting them up correctly is essential to obtaining the result you want. You also have to know what you want.
Peter
Monday, November 11, 2019
Protons (51) From peripheral cells to the brain
Hunger and satiety. Can these be modelled from a very simple energy availability concept?
This post is a minimally referenced ramble through how I see satiety working.
At the peripheral cellular level energy influx is controlled, to a large extent, via reactive oxygen species. These regulate insulin signalling which controls the translocation of GLUT4 and CD36 proteins to the cell surface and so facilitates the diffusion in to the cell of glucose and fatty acids respectively.
When a single cell has adequate calories it generates ROS, largely from the electron transport chain, which disable insulin signalling. This insulin "resistance" is there to limit excessive ingress of calories. ROS are the signal that no further calories are needed. At the most basic level cellular energy ingress is regulated by the core energy utilising apparatus of the cell. Excess substrate means excess ROS means shut down caloric ingress.
This is a self-contained cellular satiety signal of the most basic type. When the cell has fully adequate supplies of ATP, a high mitochondrial membrane voltage and a deeply reduced CoQ couple then ROS generation becomes almost inevitable.
It is a core, deep level, simple system. How it works is the subject of the Protons thread of posts, as is how it malfunctions.
I find it very, very difficult to envisage that the control of ingress of calories on a whole body basis (hunger/satiety) is not similarly integrated around an ROS generating system within dedicated neurons of the brain, with the hypothalamus being the most likely location.
I've had this as a persistent suspicion for a very long time but you can't get around to reading about everything at the same time. And I have to admit that neurological thinking about hunger and satiety has always struck me as a highly disreputable field. Insulin and satiety smell like cholesterol and the lipid hypothesis.
So yesterday I finally went and had a look for a reasonably recent review of ROS within the hypothalamus and this one came up pretty high in the PubMed listing
Impact of hypothalamic reactive oxygen species in the regulation of energy metabolism and food intake
This gives a flavour:
"Thus, it appears that NPY/AgRP neurons activation is mediated by a decrease in ROS levels while POMC neurons activation is driven by ROS (Andrews et al., 2008). Indeed, icv administration of ROS scavengers induces significantly lower c-Fos expression in POMC neurons and increases food intake during light cycle, observed via an increase of c-Fos expression in AgRP/NPY neurons (Diano et al., 2011). Similarly, addition of H2O2 depolarizes POMC neurons, increases the firing rate, and an icv injection of H2O2 causes significantly less feeding of mice after an overnight fast".
A lot of the work cited is not terribly well performed and no one has the Protons framework to slot their findings in to, but it's a start. What I am looking for is that these cell types do express GLUT proteins and CD36 proteins. I would expect them to be sampling arterial blood or CSF and integrating NADH and FADH2 inputs to their mitochondrial electron transport chain to "decide" whether there are adequate calories to consider that satiety or hunger might be the preferred descriptor of the body's current state.
Whether these cells express insulin receptors to facilitate ingress of substrate is something to be picked at. As I am completely biased against the concept that insulin is a satiety hormone, I would prefer this not to be the case but may be wrong. Time will tell. It looks logical to me that the brain would look at the nutrient levels present in excess of those being disposed of by peripheral insulin in to peripheral cells. As large numbers of peripheral cells become "full" under the influence of insulin, the brain should pick up the rising level of excess nutrients as the signal to call a halt on hunger. Doing this within the brain shouldn't need insulin, merely a set of relatively low affinity transporters to allow glucose and lipid uptake as insulin completes its peripheral function. Satiety should be picked up when enough cells have enough calories, whole body, that they no longer behave as a calorie "sump". The job of the brain is to pick up evidence from the nutrient levels that the sump is full and satiety can be declared.
For hunger, high affinity transporters would allow ROS to be generated easily and falling ROS would signal that energy availability was low.
These signals will come from the neural mitochondrial ETC generating ROS. The best ROS generating nutrients will be the most satiating. Saturated fats spring to mind if you follow the Protons thread.
Obviously there are whole load more hormones which can influence the generation of ROS within neurons. Physiology has applied layers and layers of signalling to maximise reproductive fitness. I have minimal interest in these "higher level" signals. They are there, they will modulate the core process I've been talking about but I see no way that they will do anything fundamentally different from or in opposition to the ROS system.
I'll give the rest of the review a bit of a read and see if it's worth posting about.
Peter
This post is a minimally referenced ramble through how I see satiety working.
At the peripheral cellular level energy influx is controlled, to a large extent, via reactive oxygen species. These regulate insulin signalling which controls the translocation of GLUT4 and CD36 proteins to the cell surface and so facilitates the diffusion in to the cell of glucose and fatty acids respectively.
When a single cell has adequate calories it generates ROS, largely from the electron transport chain, which disable insulin signalling. This insulin "resistance" is there to limit excessive ingress of calories. ROS are the signal that no further calories are needed. At the most basic level cellular energy ingress is regulated by the core energy utilising apparatus of the cell. Excess substrate means excess ROS means shut down caloric ingress.
This is a self-contained cellular satiety signal of the most basic type. When the cell has fully adequate supplies of ATP, a high mitochondrial membrane voltage and a deeply reduced CoQ couple then ROS generation becomes almost inevitable.
It is a core, deep level, simple system. How it works is the subject of the Protons thread of posts, as is how it malfunctions.
I find it very, very difficult to envisage that the control of ingress of calories on a whole body basis (hunger/satiety) is not similarly integrated around an ROS generating system within dedicated neurons of the brain, with the hypothalamus being the most likely location.
I've had this as a persistent suspicion for a very long time but you can't get around to reading about everything at the same time. And I have to admit that neurological thinking about hunger and satiety has always struck me as a highly disreputable field. Insulin and satiety smell like cholesterol and the lipid hypothesis.
So yesterday I finally went and had a look for a reasonably recent review of ROS within the hypothalamus and this one came up pretty high in the PubMed listing
Impact of hypothalamic reactive oxygen species in the regulation of energy metabolism and food intake
This gives a flavour:
"Thus, it appears that NPY/AgRP neurons activation is mediated by a decrease in ROS levels while POMC neurons activation is driven by ROS (Andrews et al., 2008). Indeed, icv administration of ROS scavengers induces significantly lower c-Fos expression in POMC neurons and increases food intake during light cycle, observed via an increase of c-Fos expression in AgRP/NPY neurons (Diano et al., 2011). Similarly, addition of H2O2 depolarizes POMC neurons, increases the firing rate, and an icv injection of H2O2 causes significantly less feeding of mice after an overnight fast".
A lot of the work cited is not terribly well performed and no one has the Protons framework to slot their findings in to, but it's a start. What I am looking for is that these cell types do express GLUT proteins and CD36 proteins. I would expect them to be sampling arterial blood or CSF and integrating NADH and FADH2 inputs to their mitochondrial electron transport chain to "decide" whether there are adequate calories to consider that satiety or hunger might be the preferred descriptor of the body's current state.
Whether these cells express insulin receptors to facilitate ingress of substrate is something to be picked at. As I am completely biased against the concept that insulin is a satiety hormone, I would prefer this not to be the case but may be wrong. Time will tell. It looks logical to me that the brain would look at the nutrient levels present in excess of those being disposed of by peripheral insulin in to peripheral cells. As large numbers of peripheral cells become "full" under the influence of insulin, the brain should pick up the rising level of excess nutrients as the signal to call a halt on hunger. Doing this within the brain shouldn't need insulin, merely a set of relatively low affinity transporters to allow glucose and lipid uptake as insulin completes its peripheral function. Satiety should be picked up when enough cells have enough calories, whole body, that they no longer behave as a calorie "sump". The job of the brain is to pick up evidence from the nutrient levels that the sump is full and satiety can be declared.
For hunger, high affinity transporters would allow ROS to be generated easily and falling ROS would signal that energy availability was low.
These signals will come from the neural mitochondrial ETC generating ROS. The best ROS generating nutrients will be the most satiating. Saturated fats spring to mind if you follow the Protons thread.
Obviously there are whole load more hormones which can influence the generation of ROS within neurons. Physiology has applied layers and layers of signalling to maximise reproductive fitness. I have minimal interest in these "higher level" signals. They are there, they will modulate the core process I've been talking about but I see no way that they will do anything fundamentally different from or in opposition to the ROS system.
I'll give the rest of the review a bit of a read and see if it's worth posting about.
Peter
Friday, November 08, 2019
Insulin makes you hungry (11) But not in Denmark?
Preamble. The best papers are those which challenge your ideas. When they conflict with what appears to be very hard evidence which support your mindset they become really exciting. Sometimes you just have to shrug, label the new finding as important and put it on the back burner to be ruminated about over the coming months. Sometimes a potential explanation is possible. This post is essentially a fairytale set in Denmark. It may be completely wrong. Or not. Here we go.
This paper came up in the comments to the last post from Gabor Erdosi via raphi. It is from Astrup's group in Denmark and I have to say I have a lot of time for Prof Astrup as he was one of the more influential people who objected to the gross stupidity of Denmark's transient fat tax a few years ago. The fat tax was abandoned quite rapidly as sensible EU dwelling Danes merely popped across their open border and stocked up with un-fat-taxed butter in Germany. Anyway:
The role of postprandial releases of insulin and incretin hormones in meal-induced satiety-effect of obesity and weight reduction
This is the crucial graph
Take 12 lean people, feed them a 600kcal sandwich for breakfast, wait just over three hours then offer them an ad-lib, well mixed pasta salad and see how much of this they eat.
The higher their insulin went after breakfast, the less they ate at lunchtime.
Insulin exposure is clearly associated with reduced subsequent food intake. You might be tempted to assume causality here, but you can't. It's an observational study of a very specific set of people. It can be used to generate an hypothesis, such as insulin suppresses subsequent food intake. But then you would have to test that hypothesis.
You could also simply go back through the literature to interventional studies which actually imposed changes in insulin levels and come to the opposite conclusion. Rodin et al did this here
Effect of insulin and glucose on feeding behavior
which makes that particular hypothesis untenable. Insulin makes you hungry.
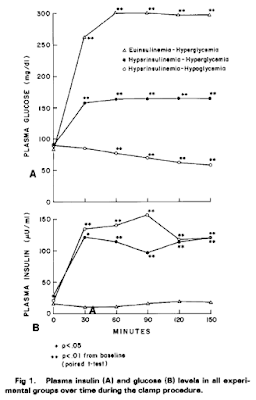
Here are the core findings, clamp values first
Here are the hunger ratings
and the amount of liquid food consumed, via a straw, through a screen:
To me personally, his study is very convincing. The principle is simple, logical and comprehensible. I would have been happier if he had also tracked FFAs in the study but we all know what insulin does to FFA levels (in the absence of fat ingestion). My personal view is that the brain looks at the availability of calories. Normoglycaemia with rapidly falling FFAs (the effect of insulin on adipocytes) is going to generate hunger. No one would expect any different. The action of insulin is the inhibition of lipolysis. Much higher levels are needed to facilitate the uptake of glucose.
We have a paradox, excellent. Direct insulin infusion makes you hungry. Insulin response to food makes you less hungry.
That is so cool.
Sooooo. Is it even remotely possible to explain the observation picked up by Prof Astrup in his 12 lean Danes? Starting from the basis that insulin drives calorie loss in to adipocytes with subsequent hunger? Speculation warning.
This group of Danes is very unusual. They have lived, on average, for 34 years in modern Denmark and they are not over weight. They have never counted a calorie, never been to Weight W@tchers, never had an eating disorder. They eat as much as they are comfortable with and eat again when they are hungry. If they pig out at Christmas they don't need to go on a diet in the New Year. They put zero effort in to being lean. That is a very special sort of person. They have normal appetite control.
When we give them a fixed calorie breakfast the insulin response varies. With these normal people I think it is a reasonable assumption that if the 600kcal is high compared to their preferred size of breakfast, the insulin level will go higher. There is more food than needed so more to store, that needs more insulin.
Members of the group who fancied many more than 600kcal for breakfast will have produced a low insulin response to the 600kcal specified by the study.
Now, it gets interesting because you cannot remotely account for a 1200kcal (3000kJ) difference in lunchtime eating by speculating about preferred breakfast size. The effect is too big.
The storage of calories is the simplest of actions of insulin. It does other things too. For these we have to go to the very, very artificial model of Veech's isolated working rat hearts. However some of the findings do have some bearing on real life.
In this paper
Substrate signaling by insulin: a ketone bodies ratio mimics insulin action in heart
we have this snippet:
"Unexpectedly, insulin increased cardiac hydraulic work but decreased net glycolytic flux and O2 consumption, improving net cardiac efficiency by 28%".
Insulin facilitates glucose diffusion in to the myocytes but partitions it in to stored glycogen. Glycolysis actually decreases but there is an increase in efficiency which gives the 28% increase in cardiac work.
Let that sink in. Insulin makes energy production more efficient while diverting calories in to storage. If you wanted to fatten someone up that seems like a good plan.
From a related paper by Veech's group
Insulin, ketone bodies, and mitochondrial energy transduction
we have a slight elaboration:
"The increase in efficiency associated with insulin administration is not readily explained by such a straightforward mechanism [as for ketones]; other factors, such as reduction of the mitochondrial NAD couple or specific effects like covalent modification of mitochondrial membrane protein, will have to be considered as possible factors altering efficiency of ATP synthesis".
Veech's model runs on glucose alone but there will undoubtedly be residual FFAs in the cardiac myocytes. You just have to wonder whether the effect of insulin is to extract these from the mitochondrial uncoupling proteins and covalently bind them in to intracellular triglycerides. An interesting idea and it would certainly tighten the coupling of the ETC.
Bottom line: Insulin, when working as it should, diverts calories to storage but increases efficiency of energy production to allow normal metabolism. I hold that this happens in the elevated insulin individuals of the lean subject group. Recall that all of these people are naturally lean. When the insulin wears off they realise, metabolically, that they have gained a (very) little weight while running a very efficient metabolism. If they have extra stored calories which, being naturally lean, they don't need, why should they eat much at lunch time?
The hypoinsulinaemic lean people get their 600kcal, decide it is way too little to bother storing and partition it towards utilisation. There is no drive towards storage, very little insulin, no insulin mediated increased efficiency. Substrate is available, it gets oxidised. Very little gets stored. This is the low insulin state. When lunch is presented to this naturally weight stable person their metabolism realises that it has not maintained fat stores after the 600kcal breakfast so they eat more at lunch time.
We have a period of high efficiency calorie conservation in the high insulin group and a period of low efficiency calorie wastage in the low insulin individuals. Because these people have that rare gem, a functional metabolism, they simply adjust subsequent food intake to reflect their previous fuel partitioning during the three hours from breakfast to lunch.
It is perfectly reasonable to mistake this scenario as an indicator that insulin is a satiety signal. Easily done. It's incorrect.
Peter
The invariable after-thought: Does insulin correlate inversely with reduced food intake in either the obese group at the start of the study or after marked weight loss?
No. Of course not.
This paper came up in the comments to the last post from Gabor Erdosi via raphi. It is from Astrup's group in Denmark and I have to say I have a lot of time for Prof Astrup as he was one of the more influential people who objected to the gross stupidity of Denmark's transient fat tax a few years ago. The fat tax was abandoned quite rapidly as sensible EU dwelling Danes merely popped across their open border and stocked up with un-fat-taxed butter in Germany. Anyway:
The role of postprandial releases of insulin and incretin hormones in meal-induced satiety-effect of obesity and weight reduction
This is the crucial graph
Take 12 lean people, feed them a 600kcal sandwich for breakfast, wait just over three hours then offer them an ad-lib, well mixed pasta salad and see how much of this they eat.
The higher their insulin went after breakfast, the less they ate at lunchtime.
Insulin exposure is clearly associated with reduced subsequent food intake. You might be tempted to assume causality here, but you can't. It's an observational study of a very specific set of people. It can be used to generate an hypothesis, such as insulin suppresses subsequent food intake. But then you would have to test that hypothesis.
You could also simply go back through the literature to interventional studies which actually imposed changes in insulin levels and come to the opposite conclusion. Rodin et al did this here
Effect of insulin and glucose on feeding behavior
which makes that particular hypothesis untenable. Insulin makes you hungry.
Here are the core findings, clamp values first
Here are the hunger ratings
and the amount of liquid food consumed, via a straw, through a screen:
To me personally, his study is very convincing. The principle is simple, logical and comprehensible. I would have been happier if he had also tracked FFAs in the study but we all know what insulin does to FFA levels (in the absence of fat ingestion). My personal view is that the brain looks at the availability of calories. Normoglycaemia with rapidly falling FFAs (the effect of insulin on adipocytes) is going to generate hunger. No one would expect any different. The action of insulin is the inhibition of lipolysis. Much higher levels are needed to facilitate the uptake of glucose.
We have a paradox, excellent. Direct insulin infusion makes you hungry. Insulin response to food makes you less hungry.
That is so cool.
Sooooo. Is it even remotely possible to explain the observation picked up by Prof Astrup in his 12 lean Danes? Starting from the basis that insulin drives calorie loss in to adipocytes with subsequent hunger? Speculation warning.
This group of Danes is very unusual. They have lived, on average, for 34 years in modern Denmark and they are not over weight. They have never counted a calorie, never been to Weight W@tchers, never had an eating disorder. They eat as much as they are comfortable with and eat again when they are hungry. If they pig out at Christmas they don't need to go on a diet in the New Year. They put zero effort in to being lean. That is a very special sort of person. They have normal appetite control.
When we give them a fixed calorie breakfast the insulin response varies. With these normal people I think it is a reasonable assumption that if the 600kcal is high compared to their preferred size of breakfast, the insulin level will go higher. There is more food than needed so more to store, that needs more insulin.
Members of the group who fancied many more than 600kcal for breakfast will have produced a low insulin response to the 600kcal specified by the study.
Now, it gets interesting because you cannot remotely account for a 1200kcal (3000kJ) difference in lunchtime eating by speculating about preferred breakfast size. The effect is too big.
The storage of calories is the simplest of actions of insulin. It does other things too. For these we have to go to the very, very artificial model of Veech's isolated working rat hearts. However some of the findings do have some bearing on real life.
In this paper
Substrate signaling by insulin: a ketone bodies ratio mimics insulin action in heart
we have this snippet:
"Unexpectedly, insulin increased cardiac hydraulic work but decreased net glycolytic flux and O2 consumption, improving net cardiac efficiency by 28%".
Insulin facilitates glucose diffusion in to the myocytes but partitions it in to stored glycogen. Glycolysis actually decreases but there is an increase in efficiency which gives the 28% increase in cardiac work.
Let that sink in. Insulin makes energy production more efficient while diverting calories in to storage. If you wanted to fatten someone up that seems like a good plan.
From a related paper by Veech's group
Insulin, ketone bodies, and mitochondrial energy transduction
we have a slight elaboration:
"The increase in efficiency associated with insulin administration is not readily explained by such a straightforward mechanism [as for ketones]; other factors, such as reduction of the mitochondrial NAD couple or specific effects like covalent modification of mitochondrial membrane protein, will have to be considered as possible factors altering efficiency of ATP synthesis".
Veech's model runs on glucose alone but there will undoubtedly be residual FFAs in the cardiac myocytes. You just have to wonder whether the effect of insulin is to extract these from the mitochondrial uncoupling proteins and covalently bind them in to intracellular triglycerides. An interesting idea and it would certainly tighten the coupling of the ETC.
Bottom line: Insulin, when working as it should, diverts calories to storage but increases efficiency of energy production to allow normal metabolism. I hold that this happens in the elevated insulin individuals of the lean subject group. Recall that all of these people are naturally lean. When the insulin wears off they realise, metabolically, that they have gained a (very) little weight while running a very efficient metabolism. If they have extra stored calories which, being naturally lean, they don't need, why should they eat much at lunch time?
The hypoinsulinaemic lean people get their 600kcal, decide it is way too little to bother storing and partition it towards utilisation. There is no drive towards storage, very little insulin, no insulin mediated increased efficiency. Substrate is available, it gets oxidised. Very little gets stored. This is the low insulin state. When lunch is presented to this naturally weight stable person their metabolism realises that it has not maintained fat stores after the 600kcal breakfast so they eat more at lunch time.
We have a period of high efficiency calorie conservation in the high insulin group and a period of low efficiency calorie wastage in the low insulin individuals. Because these people have that rare gem, a functional metabolism, they simply adjust subsequent food intake to reflect their previous fuel partitioning during the three hours from breakfast to lunch.
It is perfectly reasonable to mistake this scenario as an indicator that insulin is a satiety signal. Easily done. It's incorrect.
Peter
The invariable after-thought: Does insulin correlate inversely with reduced food intake in either the obese group at the start of the study or after marked weight loss?
No. Of course not.
Sunday, November 03, 2019
Insulin makes you hungry (10) Processing
I picked up a link to this paper from an image of a table, somewhere on Tinternet. I didn't realise which paper it was. It was really interesting from the metabolic point of view and I kept looking through the results section to find the post-prandial metabolic effect of ultra-processed foods vs unprocessed foods. You know, the effect which might determine where the calories from a given food might end up, either available for metabolism versus lost in to adipocytes.
Ultra-Processed Diets Cause Excess Calorie Intake and Weight Gain: An Inpatient Randomized Controlled Trial of Ad Libitum Food Intake
This is from the introduction
"Ultra-processed foods may facilitate overeating and the development of obesity (Poti et al., 2017) because they are typically high in calories, salt, sugar, and fat (Poti et al., 2015) and have been suggested to be engineered to have supernormal appetitive properties (Kessler, 2009; Moss, 2013; Moubarac, 2015; Schatzker, 2015). Furthermore, ultra-processed foods are theorized to disrupt gut-brain signaling and may influence food reinforcement and overall intake via mechanisms distinct from the palatability or energy density of the food (Small and DiFeliceantonio, 2019)"
What this is saying is that making foods high in calories + "supernormal appetitive properties" makes you eat more, so you become fat. And that they alter gut brain signalling to increase "reinforcement" and so increase intake. You have to put the excess calories somewhere so you get fat.
Supernormal appetitive properties. Energy density. Reinforcement.
Psychobabble.
But the bottom line is that highly processed foods do produce steady weight gain over two weeks and low processed foods do the opposite.
What we need to know is how a given food affects both the acute and chronic metabolic response, especially insulin levels and signalling. That's not going to come from the psychobabblers.
Luckily there are groups with an interest in the metabolic effects of food, rather than being lost in un-Rewarding theories of reinforcement. Like this group:
Does food insulin index in the context of mixed meals affect postprandial metabolic responses and appetite in obese adolescents with insulin resistance? A randomised cross-over trial
They compared two meals with the same low glycaemic index (GI) but differing insulin secretion indices (Insulin index, II). Both meals had very similar macros and they look suspiciously like processed vs unprocessed. Insulin (and glucose) were higher after the high II meals:
The low II meals generated less hunger in the post prandial period. I'm not sure what hunger has to do with weight gain, I get the impression from people like Hall et al that calorie intake controls weight gain. Hunger is something else, will-power dependent maybe.
The second study has some problems too. The low II food did not taste as good as the high II food. So Rewarders have a lifeline here. And if you are making a marketable commodity to be labelled as "food", anything you can do to increase its palatability is going to improve sales. If palatability should turn out to be intrinsically linked to a food's II then you are set to drive hunger combined with those increased sales. Win-win for the shareholders but not so much for the consumers. No one wants to be fat. Equally, no one can tolerate being hungry...
The second problem is that there was no increase in food intake for the high II group during the end-of-study buffet. An increase in food intake here would have been nice, to corroborate the increased hunger. But that meal was a free choice buffet being offered to people who were already obese, so poor food choices may well have over-ridden the differences in experimental meal induced hunger. Back in the days of better designed studies the post-insulin test meal was a uniform soup-like liquid, consumed via a straw through a screen so there were no visual or food choice cues to influence food intake. A pity, but there you go.
Insulin makes you hungry. Not directly, but by diverting calories in to storage. You lose those calories so your brain gets hungry. Making you fat is how insulin makes you hungry. Pure CICO but the calories-out go in to your adipocytes...
Peter
Ultra-Processed Diets Cause Excess Calorie Intake and Weight Gain: An Inpatient Randomized Controlled Trial of Ad Libitum Food Intake
This is from the introduction
"Ultra-processed foods may facilitate overeating and the development of obesity (Poti et al., 2017) because they are typically high in calories, salt, sugar, and fat (Poti et al., 2015) and have been suggested to be engineered to have supernormal appetitive properties (Kessler, 2009; Moss, 2013; Moubarac, 2015; Schatzker, 2015). Furthermore, ultra-processed foods are theorized to disrupt gut-brain signaling and may influence food reinforcement and overall intake via mechanisms distinct from the palatability or energy density of the food (Small and DiFeliceantonio, 2019)"
What this is saying is that making foods high in calories + "supernormal appetitive properties" makes you eat more, so you become fat. And that they alter gut brain signalling to increase "reinforcement" and so increase intake. You have to put the excess calories somewhere so you get fat.
Supernormal appetitive properties. Energy density. Reinforcement.
Psychobabble.
But the bottom line is that highly processed foods do produce steady weight gain over two weeks and low processed foods do the opposite.
What we need to know is how a given food affects both the acute and chronic metabolic response, especially insulin levels and signalling. That's not going to come from the psychobabblers.
Luckily there are groups with an interest in the metabolic effects of food, rather than being lost in un-Rewarding theories of reinforcement. Like this group:
Does food insulin index in the context of mixed meals affect postprandial metabolic responses and appetite in obese adolescents with insulin resistance? A randomised cross-over trial
They compared two meals with the same low glycaemic index (GI) but differing insulin secretion indices (Insulin index, II). Both meals had very similar macros and they look suspiciously like processed vs unprocessed. Insulin (and glucose) were higher after the high II meals:
The low II meals generated less hunger in the post prandial period. I'm not sure what hunger has to do with weight gain, I get the impression from people like Hall et al that calorie intake controls weight gain. Hunger is something else, will-power dependent maybe.
The second study has some problems too. The low II food did not taste as good as the high II food. So Rewarders have a lifeline here. And if you are making a marketable commodity to be labelled as "food", anything you can do to increase its palatability is going to improve sales. If palatability should turn out to be intrinsically linked to a food's II then you are set to drive hunger combined with those increased sales. Win-win for the shareholders but not so much for the consumers. No one wants to be fat. Equally, no one can tolerate being hungry...
The second problem is that there was no increase in food intake for the high II group during the end-of-study buffet. An increase in food intake here would have been nice, to corroborate the increased hunger. But that meal was a free choice buffet being offered to people who were already obese, so poor food choices may well have over-ridden the differences in experimental meal induced hunger. Back in the days of better designed studies the post-insulin test meal was a uniform soup-like liquid, consumed via a straw through a screen so there were no visual or food choice cues to influence food intake. A pity, but there you go.
Insulin makes you hungry. Not directly, but by diverting calories in to storage. You lose those calories so your brain gets hungry. Making you fat is how insulin makes you hungry. Pure CICO but the calories-out go in to your adipocytes...
Peter
Sunday, October 20, 2019
Ketogenic diets are unhelpful and dangerous for managing mitochondrial diseases. Maybe (2)
I'm hoping we all remember Sauer. If not there's a post here:
Sauer vs Lisanti
And a brief summary at the start of this one:
Sauer and 13-HODE
Sauer demonstrated that food withdrawal from rats carrying tumour xenografts makes those tumours grow like wildfire. He went on to show that it was 13-HODE which made the tumours grow and that either linoleic acid or arachidonic acid (both 13-HODE precursors) were essential for tumour growth in his model. Fasting released these carcinogenic PUFA from adipocytes when the rats were starved.
You cannot make 13-HODE without linoleic acid. Or arachidonic acid if you prefer.
Do you want to make a tumour grow? Feed your model linoleic acid. It's easy. Or you could try using a Ketocal based ketogenic diet for glioblastoma multiforme management in real live humans. There was a trial doing this recently:
Ketogenic diet treatment as adjuvant to standard treatment of glioblastoma multiforme: a feasibility and safety study
which didn't really pick up any benefit, it was only a pilot study. Serious concern about the composition of the ketogenic diet was expressed in this letter to the editor:
Problems associated with a highly artificial ketogenic diet: Letter to the Editor Re: van der Louw EJTM, Olieman JF, van den Bemt PMLA, et al. ‘Ketogenic diet treatment as adjuvant to standard treatment of glioblastoma multiforme: a feasibility and safety study’
link tweeted by Miki Ben-Dor.
Much of the diet was Ketocal based "consisting of refined vegetable oils from sunflower, soy, and palm fruit..."
This looks like an excellent source of the 13-HODE precursor linoleic acid, which Sauer might have recognised as the growth promoter in his rat models.
I would suspect that the ketogenic diet supplied benefit from ketosis, but this was largely offset by tumour promotion from the linoleic acid content.
Is there any end to the damage done by the lipid hypothesis?
Probably not.
Peter
Sauer vs Lisanti
And a brief summary at the start of this one:
Sauer and 13-HODE
Sauer demonstrated that food withdrawal from rats carrying tumour xenografts makes those tumours grow like wildfire. He went on to show that it was 13-HODE which made the tumours grow and that either linoleic acid or arachidonic acid (both 13-HODE precursors) were essential for tumour growth in his model. Fasting released these carcinogenic PUFA from adipocytes when the rats were starved.
You cannot make 13-HODE without linoleic acid. Or arachidonic acid if you prefer.
Do you want to make a tumour grow? Feed your model linoleic acid. It's easy. Or you could try using a Ketocal based ketogenic diet for glioblastoma multiforme management in real live humans. There was a trial doing this recently:
Ketogenic diet treatment as adjuvant to standard treatment of glioblastoma multiforme: a feasibility and safety study
which didn't really pick up any benefit, it was only a pilot study. Serious concern about the composition of the ketogenic diet was expressed in this letter to the editor:
Problems associated with a highly artificial ketogenic diet: Letter to the Editor Re: van der Louw EJTM, Olieman JF, van den Bemt PMLA, et al. ‘Ketogenic diet treatment as adjuvant to standard treatment of glioblastoma multiforme: a feasibility and safety study’
link tweeted by Miki Ben-Dor.
Much of the diet was Ketocal based "consisting of refined vegetable oils from sunflower, soy, and palm fruit..."
This looks like an excellent source of the 13-HODE precursor linoleic acid, which Sauer might have recognised as the growth promoter in his rat models.
I would suspect that the ketogenic diet supplied benefit from ketosis, but this was largely offset by tumour promotion from the linoleic acid content.
Is there any end to the damage done by the lipid hypothesis?
Probably not.
Peter
Friday, October 18, 2019
Total Cholesterol and major cardiovascular events
This is a pure observational study:
Low total cholesterol is associated with increased major adverse cardiovascular events in men aged ≥70 years not taking statins
As such, let's generate an hypothesis.
Let's assume total cholesterol (TC) is utterly, totally and completely meaningless. About anything.
Let's assume that TC is a surrogate for "something" (Something Good) which is really important.
Let's assume that people with high TC are doing something right. Something which the study never even thought about, let alone measured.
Let's assume that statins do absolutely nothing (being generous).
Let's assume people only get put on a statin if their TC is high, ie, they are doing Something Good.
The Something Good behaviour pattern persists despite the statin.
The statination makes the TC number decrease, which makes the statin victims look as if they are doing the Something Bad which, obviously, lowers your TC. But they're not. They do well despite the statin and despite the artificially lowered TC.
EPIC Norfolk suggests looking at HbA1c if you want to look at cardiovascular and all cause mortality. Hints about Something Good here in Norfolk?
Peter
Edit: No one (least of all me) would suggest HbA1c elevation is causative of all cause mortality. But at least it starts you looking at the correct metabolic pathways! End edit.
Low total cholesterol is associated with increased major adverse cardiovascular events in men aged ≥70 years not taking statins
As such, let's generate an hypothesis.
Let's assume total cholesterol (TC) is utterly, totally and completely meaningless. About anything.
Let's assume that TC is a surrogate for "something" (Something Good) which is really important.
Let's assume that people with high TC are doing something right. Something which the study never even thought about, let alone measured.
Let's assume that statins do absolutely nothing (being generous).
Let's assume people only get put on a statin if their TC is high, ie, they are doing Something Good.
The Something Good behaviour pattern persists despite the statin.
The statination makes the TC number decrease, which makes the statin victims look as if they are doing the Something Bad which, obviously, lowers your TC. But they're not. They do well despite the statin and despite the artificially lowered TC.
EPIC Norfolk suggests looking at HbA1c if you want to look at cardiovascular and all cause mortality. Hints about Something Good here in Norfolk?
Peter
Edit: No one (least of all me) would suggest HbA1c elevation is causative of all cause mortality. But at least it starts you looking at the correct metabolic pathways! End edit.
Saturday, October 12, 2019
Metformin (11) a SHORT paradox
I'll just throw this one out there as I found it while looking for something else:
Metformin paradoxically worsens insulin resistance in SHORT syndrome
SHORT syndrome is a (very, very rare) genetic failure of insulin signalling at the PIK3 regulatory subunit 1 level. Insulin binds to its receptor but signalling fails due to a single downstream gene defect giving severe insulin resistance. People with this syndrome are, needless to say, very thin. They maintain normoglycaemia using a very high level of insulin which does, given a high enough concentration, produce normoglycaemia. There doesn't appear to be any problem with secreting insulin from the pancreas. In fact, to overcome the failure of insulin signalling during her OGTT the patient's pancreas produced a plasma insulin of 688mIU/l*. In new money that is just under 5000pmol. As in roughly ten times what you might expect. Severe, but not quite insuperable, insulin resistance.
*The paper specifies insulin in mIU/ml. I'm assuming this is a typo or a font failure because clinical insulin concentrations are usually expressed as microIU/ml or mIU/l. Obviously if it really is 688mIU/ml the concentrations will be 1000 times those quoted. Gulp. People really should use the SI system.
Back to the patient. The obvious thing to do is to give an insulin sensitising agent, number one in popularity nowadays being metformin.
This turned out to be a bit of a boo boo.
During an OGTT under metformin the patient's insulin resistance worsened and mild hyperglycaemia ensured but this was despite a plasma insulin concentration which was simply too high to measure. The lab could measure up to around 7000pmol (pax typos) and it looks like the curve went MUCH higher than that.
That is despite metformin's predictable and recently found ability to suppress insulin release from mouse islets.
From the Protons perspective metformin blunts insulin signalling via blockade of mitochondrial glycerol-3-phosphate dehydrogenase. Its beneficial effects to increase insulin sensitivity come from reduced exposure to insulin signalling in peripheral cells. The peripheral cells of a person with SHORT syndrome barely see insulin signalling at all even without the metformin. You would hardly expect further blocking any residual insulin signalling to help matters. It doesn't.
Metformin paradoxically worsens insulin resistance in SHORT syndrome
SHORT syndrome is a (very, very rare) genetic failure of insulin signalling at the PIK3 regulatory subunit 1 level. Insulin binds to its receptor but signalling fails due to a single downstream gene defect giving severe insulin resistance. People with this syndrome are, needless to say, very thin. They maintain normoglycaemia using a very high level of insulin which does, given a high enough concentration, produce normoglycaemia. There doesn't appear to be any problem with secreting insulin from the pancreas. In fact, to overcome the failure of insulin signalling during her OGTT the patient's pancreas produced a plasma insulin of 688mIU/l*. In new money that is just under 5000pmol. As in roughly ten times what you might expect. Severe, but not quite insuperable, insulin resistance.
*The paper specifies insulin in mIU/ml. I'm assuming this is a typo or a font failure because clinical insulin concentrations are usually expressed as microIU/ml or mIU/l. Obviously if it really is 688mIU/ml the concentrations will be 1000 times those quoted. Gulp. People really should use the SI system.
Back to the patient. The obvious thing to do is to give an insulin sensitising agent, number one in popularity nowadays being metformin.
This turned out to be a bit of a boo boo.
During an OGTT under metformin the patient's insulin resistance worsened and mild hyperglycaemia ensured but this was despite a plasma insulin concentration which was simply too high to measure. The lab could measure up to around 7000pmol (pax typos) and it looks like the curve went MUCH higher than that.
That is despite metformin's predictable and recently found ability to suppress insulin release from mouse islets.
From the Protons perspective metformin blunts insulin signalling via blockade of mitochondrial glycerol-3-phosphate dehydrogenase. Its beneficial effects to increase insulin sensitivity come from reduced exposure to insulin signalling in peripheral cells. The peripheral cells of a person with SHORT syndrome barely see insulin signalling at all even without the metformin. You would hardly expect further blocking any residual insulin signalling to help matters. It doesn't.
EDIT in 2025: Metformin does NOT sensitise to insulin at all. The improvements in insulin numbers are secondary to the non hormonal suppression of hepatic glucose output. Obviously any weight loss drug MUST be suppressing either insulin levels per se or the resultant insulin signalling. END EDIT.
It's the sort of paradox which only happens when you are in the wrong paradigm of metformin's mechanism of action. Might have been a chance to make progress...
Peter
Addendum: The lady in question did not seem to enjoy her experience with the medics too much:
"As we intended to check the effects of this approach, an extended 75 g OGTT was performed on metformin 4 days later. This showed dramatic and paradoxical worsening of insulin tolerance with insulin concentrations above the upper assay detection limit (Fig. 1b). Metformin treatment was discontinued. She was discharged home on Dydrogesterone and vitamin D supplementation. We planned to perform investigations on other family members, and particularly on her younger brother, but despite several reminders they failed to attend clinic appointments as well as declined admission to the hospital".
Peter
Addendum: The lady in question did not seem to enjoy her experience with the medics too much:
"As we intended to check the effects of this approach, an extended 75 g OGTT was performed on metformin 4 days later. This showed dramatic and paradoxical worsening of insulin tolerance with insulin concentrations above the upper assay detection limit (Fig. 1b). Metformin treatment was discontinued. She was discharged home on Dydrogesterone and vitamin D supplementation. We planned to perform investigations on other family members, and particularly on her younger brother, but despite several reminders they failed to attend clinic appointments as well as declined admission to the hospital".
Friday, October 11, 2019
Ketogenic diets are unhelpful and dangerous for managing mitochondrial diseases. Maybe.
I think this is probably an abstract from a short communication at a conference. I picked it up from Miki Ben-Dor on twitter

I think we can say that ketogenic diets are pretty rubbish for managing mitochondrial illnesses.
By chance there is also a twitter discussion on-going relating to omega 6 based ketogenic diets.
First, nutritionists LOVE omega 6 PUFA and HATE saturated fats. In case anyone hadn't noticed. This tweet came from laura cooper, this time picked up via Raphi and Tucker.
Supported by this
It seems very likely that we can combine these two concepts and come up with some sort of an explanation.
I think we can accept that the ketone induced metabolic changes noted by Veech in isolated working rat hearts, resulting in increased energy yield per ATP molecule, still apply even with poorly functional mitochondria, because there is comparable improvement in the control of abnormal mitochondria induced intractable epilepsy vs ordinary intractable epilepsy. The ketogenic diet is clearly doing something...
That's good.
Everything else is bad.
If there is a significant problem with the structure/function of complex I ketones will not be directly helpful. They deliver acetyl-CoA to the TCA and essentially nothing else. There will be a small FADH2 input from succinate dehydrogenase but all other electrons will be presented as NADH, which needs a functional complex I to do anything much.
To bypass a poorly functional complex I we really need input as FADH2 directly to the CoQ couple without having to turn the TCA. That means beta oxidation of fatty acids, in particular it needs those fatty acids to be fully saturated because electron transporting flavoprotein only receives electrons to form FADH2 from the first desaturation step at the start of beta oxidation. Any double bonds skip this step.
Using PUFA immediately reduces energy sourced via this route.
The next thing we need to realise that modern nutritionist derived ketogenic diets cause, amongst other things, pancreatitis. I posted about pancreatitis, Intralipid and propofol here. It should come as no surprise that the side effects (from here) of PUFA based ketogenic diets in children can be severe, they're probably a lot higher in PUFA than even F3666 rodent chow...
"Other early-onset complications, in order of frequency, were hypertriglyceridemia, transient hyperuricemia, hypercholesterolemia, various infectious diseases, symptomatic hypoglycemia, hypoproteinemia, hypomagnesemia, repetitive hyponatremia, low concentrations of high-density lipoprotein, lipoid pneumonia due to aspiration, hepatitis, acute pancreatitis, and persistent metabolic acidosis. Late-onset complications also included osteopenia, renal stones, cardiomyopathy, secondary hypocarnitinemia, and iron-deficiency anemia".
Then there are cardiolipins. Each cytochrome C molecule is anchored to the outer surface of the inner mitochondrial membrane by four lipid anchors. Their nature is largely controlled by the dietary lipid supply. Modern PUFA based ketogenic diets will result in highly unsaturated cardiolipin anchors. Damaged mitochondria produce an excess of ROS. ROS break PUFA based cardiolipins giving apoptosis or, if ATP levels are too low for this, necrosis. Not going to do your ragged red muscle fibres any good. Or you cardiomyopathic cardiomyocytes.
I could go on, but you get the flavour.
Is there any end to the damage done by the lipid hypothesis?
Probably not.
Peter

I think we can say that ketogenic diets are pretty rubbish for managing mitochondrial illnesses.
By chance there is also a twitter discussion on-going relating to omega 6 based ketogenic diets.
First, nutritionists LOVE omega 6 PUFA and HATE saturated fats. In case anyone hadn't noticed. This tweet came from laura cooper, this time picked up via Raphi and Tucker.
Supported by this
It seems very likely that we can combine these two concepts and come up with some sort of an explanation.
I think we can accept that the ketone induced metabolic changes noted by Veech in isolated working rat hearts, resulting in increased energy yield per ATP molecule, still apply even with poorly functional mitochondria, because there is comparable improvement in the control of abnormal mitochondria induced intractable epilepsy vs ordinary intractable epilepsy. The ketogenic diet is clearly doing something...
That's good.
Everything else is bad.
If there is a significant problem with the structure/function of complex I ketones will not be directly helpful. They deliver acetyl-CoA to the TCA and essentially nothing else. There will be a small FADH2 input from succinate dehydrogenase but all other electrons will be presented as NADH, which needs a functional complex I to do anything much.
To bypass a poorly functional complex I we really need input as FADH2 directly to the CoQ couple without having to turn the TCA. That means beta oxidation of fatty acids, in particular it needs those fatty acids to be fully saturated because electron transporting flavoprotein only receives electrons to form FADH2 from the first desaturation step at the start of beta oxidation. Any double bonds skip this step.
Using PUFA immediately reduces energy sourced via this route.
The next thing we need to realise that modern nutritionist derived ketogenic diets cause, amongst other things, pancreatitis. I posted about pancreatitis, Intralipid and propofol here. It should come as no surprise that the side effects (from here) of PUFA based ketogenic diets in children can be severe, they're probably a lot higher in PUFA than even F3666 rodent chow...
"Other early-onset complications, in order of frequency, were hypertriglyceridemia, transient hyperuricemia, hypercholesterolemia, various infectious diseases, symptomatic hypoglycemia, hypoproteinemia, hypomagnesemia, repetitive hyponatremia, low concentrations of high-density lipoprotein, lipoid pneumonia due to aspiration, hepatitis, acute pancreatitis, and persistent metabolic acidosis. Late-onset complications also included osteopenia, renal stones, cardiomyopathy, secondary hypocarnitinemia, and iron-deficiency anemia".
Then there are cardiolipins. Each cytochrome C molecule is anchored to the outer surface of the inner mitochondrial membrane by four lipid anchors. Their nature is largely controlled by the dietary lipid supply. Modern PUFA based ketogenic diets will result in highly unsaturated cardiolipin anchors. Damaged mitochondria produce an excess of ROS. ROS break PUFA based cardiolipins giving apoptosis or, if ATP levels are too low for this, necrosis. Not going to do your ragged red muscle fibres any good. Or you cardiomyopathic cardiomyocytes.
I could go on, but you get the flavour.
Is there any end to the damage done by the lipid hypothesis?
Probably not.
Peter
Saturday, October 05, 2019
Confirmation bias is so nice
Excellent work:
Scientific discourse in the era of open science: a response to Hall et al. regarding the Carbohydrate-Insulin Model
Via raphi on twitter
Peter
Scientific discourse in the era of open science: a response to Hall et al. regarding the Carbohydrate-Insulin Model
Via raphi on twitter
Peter
Tuesday, October 01, 2019
Personal update 2019
Okay, personal update time.
Back in the middle of May this year Paul emailed me to let me know that Dr Kwasniewski had died at the age of 82. The possibility of his having had bowel cancer several years ago is apparently nearly impossible to follow up on but it doesn't appear to have been directly related to his passing away. I'd been meaning to post on this but never quite got round to it until Marco also emailed me with the same news last week. The Optimal Diet (OD) has served me well for about 17 years or so.
May was an interesting time for me. For a set of reasons not at all related to my own health I had been tempted to try the scenario of a paleolithic ketogenic diet, much along the lines of the Paleomedicina PKD protocol. I was basically interested in the level of practicality involved before suggesting it to a friend with a "modern-ketosis" resistant neurological problem. The practicalities eventually proved too problematic so the PKD option was never taken up.
I personally never expected that the PKD would change much for me.
I was wrong.
First, I stopped snoring. As far as I am aware, completely. Within a few days. I have a severely deformed nasal septum, probably traumatic in origin (if playing "toss the caber" as a kid with a larch pole, don't throw it straight up in the air in case it comes vertically back downwards directly on to your nose. Ouch). Both nostrils are severely narrowed. I never expected to ever stop snoring.
Second, the low back pain from which I got enormous relief with the OD, went. I've had three minor positional back injuries in five months but each resolution has been incredibly rapid with minimal analgesic needs.
My minor dry skin problems went within a few days, though this coincided with onset of decent access to sunlight in May, the Spring had been cool here in the UK.
Oh, and I dropped from 66kg to 62kg in a month, 11-12% body fat to 9%, estimated by lower body impedance on a set of Tanita home scales.
I carried on with the PKD.
So now I am stuck.
I really enjoy not being awakened by my wife to get me to roll back to sleeping on my side again, sometimes several times a night. I like having no back pain. I like the continued muscle strength development at the bouldering wall.
On the downside it is very socially uncomfortable. It has really brought home to me how utterly easy standard modern ketogenic eating really is. A bit of cooking and a few sweeteners and there is almost nothing you can't have within the diet.
Over the months on PKD I've added in very occasional cheese and a very, very occasional glass of Proseco on a Friday night, without apparent problems. Adding cauliflower or broccoli triggered low back soreness (I have to wonder if this is a nocebo effect, not exactly double blind!).
So nowadays I'm thinking about protein, GH, IGF-1 and insulin. I've always been cautious about protein levels but there are features about higher protein within a solidly ketogenic background that limits IGF-1 generation per unit GH secretion.
There are a number of posts there.
Currently I am, somewhat reluctantly, almost completely plant free. I'm no guru on this way of eating any more than on anything else, plus I'm very late to the party!
Peter
Back in the middle of May this year Paul emailed me to let me know that Dr Kwasniewski had died at the age of 82. The possibility of his having had bowel cancer several years ago is apparently nearly impossible to follow up on but it doesn't appear to have been directly related to his passing away. I'd been meaning to post on this but never quite got round to it until Marco also emailed me with the same news last week. The Optimal Diet (OD) has served me well for about 17 years or so.
May was an interesting time for me. For a set of reasons not at all related to my own health I had been tempted to try the scenario of a paleolithic ketogenic diet, much along the lines of the Paleomedicina PKD protocol. I was basically interested in the level of practicality involved before suggesting it to a friend with a "modern-ketosis" resistant neurological problem. The practicalities eventually proved too problematic so the PKD option was never taken up.
I personally never expected that the PKD would change much for me.
I was wrong.
First, I stopped snoring. As far as I am aware, completely. Within a few days. I have a severely deformed nasal septum, probably traumatic in origin (if playing "toss the caber" as a kid with a larch pole, don't throw it straight up in the air in case it comes vertically back downwards directly on to your nose. Ouch). Both nostrils are severely narrowed. I never expected to ever stop snoring.
Second, the low back pain from which I got enormous relief with the OD, went. I've had three minor positional back injuries in five months but each resolution has been incredibly rapid with minimal analgesic needs.
My minor dry skin problems went within a few days, though this coincided with onset of decent access to sunlight in May, the Spring had been cool here in the UK.
Oh, and I dropped from 66kg to 62kg in a month, 11-12% body fat to 9%, estimated by lower body impedance on a set of Tanita home scales.
I carried on with the PKD.
So now I am stuck.
I really enjoy not being awakened by my wife to get me to roll back to sleeping on my side again, sometimes several times a night. I like having no back pain. I like the continued muscle strength development at the bouldering wall.
On the downside it is very socially uncomfortable. It has really brought home to me how utterly easy standard modern ketogenic eating really is. A bit of cooking and a few sweeteners and there is almost nothing you can't have within the diet.
Over the months on PKD I've added in very occasional cheese and a very, very occasional glass of Proseco on a Friday night, without apparent problems. Adding cauliflower or broccoli triggered low back soreness (I have to wonder if this is a nocebo effect, not exactly double blind!).
So nowadays I'm thinking about protein, GH, IGF-1 and insulin. I've always been cautious about protein levels but there are features about higher protein within a solidly ketogenic background that limits IGF-1 generation per unit GH secretion.
There are a number of posts there.
Currently I am, somewhat reluctantly, almost completely plant free. I'm no guru on this way of eating any more than on anything else, plus I'm very late to the party!
Peter
Monday, September 23, 2019
The paradoxical fat mice (2)
This is the intra-peritoneal insulin tolerance test (ITT) result from the mice in
Caloric Restriction Paradoxically Increases Adiposity in Mice With Genetically Reduced Insulin
as mentioned a post or two ago and which needs some sort of an explanation:
The two asterisks denote that for both of the calorie restricted groups of mice there is an elevated glucose compared to the ad-lib groups in the late part of the ITT, irrespective of whether the insulin gene dose had been reduced by 50% or 75%. Obviously the effect is biologically trivial but the p value of less than 0.05 makes me think the effect is real.
I think to understand this we have to go back some time and look at the concept that metformin has no effect on blood glucose in the absence of insulin. This is the graph from here, discussed here:

My interpretation was/is that, on a background of no metformin (upper line) that insulin (given at 90 mins) generated insulin induced insulin resistance from about sixty minutes later (time 150 mins) and this become p less than 0.05 by 90 minutes after the insulin (time 180 mins), illustrated by the failure to generate insulin-induced insulin resistance in the metformin treated mice (the lower trace).
This is insulin-induced insulin resistance in type 1 diabetic mice revealed by metformin treatment.
Next we can look at type 1 diabetic mice chronically treated with long acting exogenous insulin for a few weeks before an ITT. These have pre existing insulin-induced insulin resistance before any intra-peritoneal short acting insulin is given, taken from here, previously discussed here:
In this case the mice with established insulin resistance simply developed hyperglycaemia when injected with intra-peritoneal short acting insulin. There are no p values but by eyeball the 120 minute value on the upper curve looks like it might be stastically significantly elevated compared to time zero. The message here is that insulin given to an insulin resistance patient can produce hyperglycaemia.
This is the Somogyi Effect. It is real.
In the real world insulin secretion and insulin sensitivity are carefully balanced. Anything which increases insulin secretion increases tissue exposure to insulin and down regulates insulin action. Insulin is the messenger between insulin secretion at the pancreas and insulin response/resistance at the tissues. This is in addition to the shared use of reactive oxygen species to generate both insulin secretion and insulin responsiveness.
We know that the mice with full Ins2 knockout and with or without Ins1 partial knockout are phenotypically normal and have fairly normal insulin levels in their blood.
Aside: To actually get reduced insulin levels Johnson's lab have more recently used a full Ins1 knockout with or without partial Ins2 knockout. The partial Ins2 knockouts do have lowered insulin, are slim, don't get fatty liver and live much longer than they should do. It's all in here. Nice. End aside.
So reduced insulin genotype mice should be more insulin sensitive than full insulin complement mice, though we didn't have a fully normal group in either of the papers from the Johnson lab.
Without calorie restriction (CR) Ins1 partial knockout have their insulin system in balance. With caloric restriction they are so insulin sensitive that when they have an insulogenic calorie restricted small meal they lose calories in to their adipocytes and enter torpor, ie insulin signalling is verging on pathological. So they get fat too. They are not insulin resistant.
But during an ITT they do not merely have the modest insulin levels they might produce in response to their normal small meals. They get 0.75iu/kg of insulin IP, probably more than they have ever seen before. It works. Insulin signalling drops plasma glucose for about 30 minutes. At this point the tissues realise that they are seeing more insulin than they have ever seen in their lives. Insulin-induced insulin resistance kicks in and with it the Somogyi Effect to give elevated glucose.
I think the graph at the top of this page shows an acute onset insulin-induced insulin resistance.
This insulin resistance effect appears to be releasing glucose from the liver, or failing to oppose glucagon action here. It might also have allowed a release of free fatty acids from those greedy adipocytes which precipitated daily torpor. It is just possible that the transient insulin resistant state during the brief ITT might be the only few hours of the entire life of the CR mice that they were not hungry...
Peter
Caloric Restriction Paradoxically Increases Adiposity in Mice With Genetically Reduced Insulin
as mentioned a post or two ago and which needs some sort of an explanation:
The two asterisks denote that for both of the calorie restricted groups of mice there is an elevated glucose compared to the ad-lib groups in the late part of the ITT, irrespective of whether the insulin gene dose had been reduced by 50% or 75%. Obviously the effect is biologically trivial but the p value of less than 0.05 makes me think the effect is real.
I think to understand this we have to go back some time and look at the concept that metformin has no effect on blood glucose in the absence of insulin. This is the graph from here, discussed here:

My interpretation was/is that, on a background of no metformin (upper line) that insulin (given at 90 mins) generated insulin induced insulin resistance from about sixty minutes later (time 150 mins) and this become p less than 0.05 by 90 minutes after the insulin (time 180 mins), illustrated by the failure to generate insulin-induced insulin resistance in the metformin treated mice (the lower trace).
This is insulin-induced insulin resistance in type 1 diabetic mice revealed by metformin treatment.
Next we can look at type 1 diabetic mice chronically treated with long acting exogenous insulin for a few weeks before an ITT. These have pre existing insulin-induced insulin resistance before any intra-peritoneal short acting insulin is given, taken from here, previously discussed here:
In this case the mice with established insulin resistance simply developed hyperglycaemia when injected with intra-peritoneal short acting insulin. There are no p values but by eyeball the 120 minute value on the upper curve looks like it might be stastically significantly elevated compared to time zero. The message here is that insulin given to an insulin resistance patient can produce hyperglycaemia.
This is the Somogyi Effect. It is real.
In the real world insulin secretion and insulin sensitivity are carefully balanced. Anything which increases insulin secretion increases tissue exposure to insulin and down regulates insulin action. Insulin is the messenger between insulin secretion at the pancreas and insulin response/resistance at the tissues. This is in addition to the shared use of reactive oxygen species to generate both insulin secretion and insulin responsiveness.
We know that the mice with full Ins2 knockout and with or without Ins1 partial knockout are phenotypically normal and have fairly normal insulin levels in their blood.
Aside: To actually get reduced insulin levels Johnson's lab have more recently used a full Ins1 knockout with or without partial Ins2 knockout. The partial Ins2 knockouts do have lowered insulin, are slim, don't get fatty liver and live much longer than they should do. It's all in here. Nice. End aside.
So reduced insulin genotype mice should be more insulin sensitive than full insulin complement mice, though we didn't have a fully normal group in either of the papers from the Johnson lab.
Without calorie restriction (CR) Ins1 partial knockout have their insulin system in balance. With caloric restriction they are so insulin sensitive that when they have an insulogenic calorie restricted small meal they lose calories in to their adipocytes and enter torpor, ie insulin signalling is verging on pathological. So they get fat too. They are not insulin resistant.
But during an ITT they do not merely have the modest insulin levels they might produce in response to their normal small meals. They get 0.75iu/kg of insulin IP, probably more than they have ever seen before. It works. Insulin signalling drops plasma glucose for about 30 minutes. At this point the tissues realise that they are seeing more insulin than they have ever seen in their lives. Insulin-induced insulin resistance kicks in and with it the Somogyi Effect to give elevated glucose.
I think the graph at the top of this page shows an acute onset insulin-induced insulin resistance.
This insulin resistance effect appears to be releasing glucose from the liver, or failing to oppose glucagon action here. It might also have allowed a release of free fatty acids from those greedy adipocytes which precipitated daily torpor. It is just possible that the transient insulin resistant state during the brief ITT might be the only few hours of the entire life of the CR mice that they were not hungry...
Peter
Saturday, September 21, 2019
Ketones in Tehran
Just a one-liner
This is a quite fascinating paper from Iran. Bear in mind that none of the authors appears to be a native english speaker and that they could really have done with some editorial assistance, but the results seem quite significant to me. Seyfried gets a thank you for input to the study design but clearly was not an author.
Feasibility, Safety, and Beneficial Effects of MCT Based Ketogenic Diet for Breast Cancer Treatment: A Randomized Controlled Trial Study
They recruited patients who were deemed to need chemotherapy before breast cancer surgery. Half got chemo alone and the other half got chemo plus 12 weeks on a calorie restricted, MCT based ketogenic diet. They all went to surgery and were followed up post-op for about 30 months.
As far as I can make out 30 people in each group completed the intervention. Each of the cross ticks ("censored") are patients lost to follow up in some way, around 10 in each group.
It looks very much like none of the available for follow up patients in the intervention group died. Forty percent (about ten people?) died in the control group, p=0.04.
This is from a 12 week ketogenic pre-surgery intervention. If a drug had produced this effect it would be a blockbuster. There was no instruction to stay ketogenic after the 12 week trial period finished, though there are hints that at least some of the women did.
Interesting, to say the least.
Peter
This is a quite fascinating paper from Iran. Bear in mind that none of the authors appears to be a native english speaker and that they could really have done with some editorial assistance, but the results seem quite significant to me. Seyfried gets a thank you for input to the study design but clearly was not an author.
Feasibility, Safety, and Beneficial Effects of MCT Based Ketogenic Diet for Breast Cancer Treatment: A Randomized Controlled Trial Study
They recruited patients who were deemed to need chemotherapy before breast cancer surgery. Half got chemo alone and the other half got chemo plus 12 weeks on a calorie restricted, MCT based ketogenic diet. They all went to surgery and were followed up post-op for about 30 months.
As far as I can make out 30 people in each group completed the intervention. Each of the cross ticks ("censored") are patients lost to follow up in some way, around 10 in each group.
It looks very much like none of the available for follow up patients in the intervention group died. Forty percent (about ten people?) died in the control group, p=0.04.
This is from a 12 week ketogenic pre-surgery intervention. If a drug had produced this effect it would be a blockbuster. There was no instruction to stay ketogenic after the 12 week trial period finished, though there are hints that at least some of the women did.
Interesting, to say the least.
Peter
Sunday, September 15, 2019
The paradoxical fat mice (1)
This paper is very interesting. I think I picked it up via Raphi on twitter. It comes from Jim Johnson's lab.
Caloric Restriction Paradoxically Increases Adiposity in Mice With Genetically Reduced Insulin
The background is in these two papers:
Phenotypic alterations in insulin-deficient mutant mice
Compensatory Responses in Mice Carrying a Null Mutation for Ins1 or Ins2
The paradoxical mice all had the Ins2 gene fully knocked out and in addition to this some mice also had one allele for the Ins1 gene knocked out (Ins1+/-). So the mice in the study had either a half or a quarter of the normal mouse insulin gene complement. Some mice were fed ad-lib, some were 40% calorie restricted (CR).
The CR, lowest insulin gene group (Ins1+/-) had significantly elevated total fat mass and a significantly elevated percentage of bodyweight as fat. That's a paradox to the insulin hypothesis of obesity and so really interesting. The Ins1+/+ group also had a (ns) increase in percentage body fat but not in absolute fat mass, so the trend is there too, but only a trend.
Metabolically, the split is between ad-lib vs CR groups.
All mice had the same maximal insulin response to a 2g/kg intraperitoneal glucose tolerance test but the CR groups had a very significantly reduced peak and AUC for glucose, ie they were much more insulin sensitive. The intra-peritoneal insulin tolerance test result might be worth a post in its own right, it's paradoxical too but there won't be space to cover it today.
So let's have a look at energy expenditure (EE) from Fig3 C.
To make things a bit clearer I've copy pasted the light period from the left half of the graph on to the end of the dark period to give more of an idea of the EE curves are really like during dark to light transition:
The red line starts horizontally with no significant difference in EE between ad-lib fed mice or CR mice. There is a modest increase during the dark (active) period when the CR mice get their three very small meals, as indicated. After the last meal a precipitous and highly significant fall in EE occurs. The mice enter torpor, a state of extreme lassitude and hypothermia. At around two hours in to the next light period the mice wake up and EE returns to just below that of the ad-lib mice and the cycle repeats. The ad-lib mice behave like normal mice.
The CR mice have a profound hypometabolic period every day. You could argue, if you are a cico-tard, that this is why they store excess fat. They eat all the food they can get but expend relatively little energy so they become fat: CICO. But I would disagree.
Here's my guess as to what is happening. Speculation warning.
We know that the CR mice are exquisitely insulin sensitive. They are that way because they have a low number of insulin genes and they never get enough food to trigger a major insulin spike anyway. The CR is the dominant factor but it needs the genetic background to get the paradox to occur. Insulin-induced insulin resistance, acute or chronic, does not occur due to lifetime low insulin exposure. The fact that all mice are capable of producing the same maximal insulin response to an IPGTT does not mean that the CR group experience an equivalent insulin exposure to the ad-lib group during their routine lives. They never get enough food to trigger a maximal insulin response.
The CR mice spent the bulk of the light period with a slightly low EE. Dark period arrives and with it food. As the food is eaten there is an upward trend in EE followed by a drop. The second small meal arrives, again an upswing followed by a drop. The third and final meal gives the same upswing but the drop in EE which follows just goes on downward. The mice enter torpor, a state of profound lassitude and hypothermia.
I think torpor happens because the mice simply have no accessible calories.
This is despite the fact that it occurs immediately after the third of their calorie restricted meals. Their problem is that the meals generate an insulin response. The mice are so insulin sensitive that calories are lost in to adipocytes (and probably hepatocytes) under the over-effective action of insulin.
They lose calories in to adipocytes. These are calories out. The adipocytes get bigger with the lost fat.
Torpor occurs BECAUSE the mice have become fatter.
This is the equivalent of the hunger which follows for a human under a euglycaemic (or even hyperglycaemic) hyperinsulinaemic clamp. There is no hypoglycaemia but fatty acids become locked in to adipocytes by the hyperisulinaemia and hunger follows due a lack of available calories. I posted about it here.
At two hours in to the light period insulin drops low enough to allow lipolysis. The mice wake up.
That's all.
Except: Why do the CR mice have paradoxically (although ns) elevated fasting insulin cf the ad lib mice? There are two reasons. Here are the blood sample times added to the EE graph. The arrows are not quite in the correct clock times, as detailed in the methods, but the times related to feeding/metabolism are approximately correct.
The green arrow is the sampling time for the ad-lib fed mice. It is about six hours in to the light period and the mice would normally have been asleep during the hours leading up to it. Light-period snacking, from the respiratory exchange ratio (RER) graph in Fig3 D, would not normally have started by this time so it's a very simple physiological fasting sample.
The blue arrow for the fasting CR mice just hits the end of torpor. I'm not sure these mice ever have a time when they wouldn't eat, given the chance, but here they are in their hypometabolic phase and have minimal access to calories. At this time insulin is actually a little (ns) higher than for the ad-lib groups (Fig1 D). Higher insulin means fat stays in adipocytes. Why is insulin high?
Calorie restriction does many things in addition to dropping metabolic rate. If you fast a hard working group of humans for 5 days they develop a post prandial increase in GIP (glucose-dependent insulinotropic hormone). This was found in the CR mice in both the fasting and fed state (Fig4 C). GIP facilitates insulin release, hence insulin is a little higher the CR mice and loss of calories in to adipocytes more severe, necessitating torpor.
It's interesting as to why GIP might be elevated under hunger conditions. Possibly generating and saving fat becomes a priority when calories are low. The Ins1+/- CR mice certainly have the highest RER (>1.05) after their third meal, suggesting that they prioritise the conversion of glucose to fat. This DNL, should it occur in the liver, might go some way to explaining the elevated triglycerides in both of the CR groups. Maybe. Accentuated DNL in people who have undergone massive weight loss via gastric bypass surgery is routine during an OGTT. Like these people.
Anyway, I'll stop now. This post is about 1/4 the length it started out as, so if corners seem a bit cut then mea culpa.
Peter
Caloric Restriction Paradoxically Increases Adiposity in Mice With Genetically Reduced Insulin
The background is in these two papers:
Phenotypic alterations in insulin-deficient mutant mice
Compensatory Responses in Mice Carrying a Null Mutation for Ins1 or Ins2
The paradoxical mice all had the Ins2 gene fully knocked out and in addition to this some mice also had one allele for the Ins1 gene knocked out (Ins1+/-). So the mice in the study had either a half or a quarter of the normal mouse insulin gene complement. Some mice were fed ad-lib, some were 40% calorie restricted (CR).
The CR, lowest insulin gene group (Ins1+/-) had significantly elevated total fat mass and a significantly elevated percentage of bodyweight as fat. That's a paradox to the insulin hypothesis of obesity and so really interesting. The Ins1+/+ group also had a (ns) increase in percentage body fat but not in absolute fat mass, so the trend is there too, but only a trend.
Metabolically, the split is between ad-lib vs CR groups.
All mice had the same maximal insulin response to a 2g/kg intraperitoneal glucose tolerance test but the CR groups had a very significantly reduced peak and AUC for glucose, ie they were much more insulin sensitive. The intra-peritoneal insulin tolerance test result might be worth a post in its own right, it's paradoxical too but there won't be space to cover it today.
So let's have a look at energy expenditure (EE) from Fig3 C.
To make things a bit clearer I've copy pasted the light period from the left half of the graph on to the end of the dark period to give more of an idea of the EE curves are really like during dark to light transition:
The red line starts horizontally with no significant difference in EE between ad-lib fed mice or CR mice. There is a modest increase during the dark (active) period when the CR mice get their three very small meals, as indicated. After the last meal a precipitous and highly significant fall in EE occurs. The mice enter torpor, a state of extreme lassitude and hypothermia. At around two hours in to the next light period the mice wake up and EE returns to just below that of the ad-lib mice and the cycle repeats. The ad-lib mice behave like normal mice.
The CR mice have a profound hypometabolic period every day. You could argue, if you are a cico-tard, that this is why they store excess fat. They eat all the food they can get but expend relatively little energy so they become fat: CICO. But I would disagree.
Here's my guess as to what is happening. Speculation warning.
We know that the CR mice are exquisitely insulin sensitive. They are that way because they have a low number of insulin genes and they never get enough food to trigger a major insulin spike anyway. The CR is the dominant factor but it needs the genetic background to get the paradox to occur. Insulin-induced insulin resistance, acute or chronic, does not occur due to lifetime low insulin exposure. The fact that all mice are capable of producing the same maximal insulin response to an IPGTT does not mean that the CR group experience an equivalent insulin exposure to the ad-lib group during their routine lives. They never get enough food to trigger a maximal insulin response.
The CR mice spent the bulk of the light period with a slightly low EE. Dark period arrives and with it food. As the food is eaten there is an upward trend in EE followed by a drop. The second small meal arrives, again an upswing followed by a drop. The third and final meal gives the same upswing but the drop in EE which follows just goes on downward. The mice enter torpor, a state of profound lassitude and hypothermia.
I think torpor happens because the mice simply have no accessible calories.
This is despite the fact that it occurs immediately after the third of their calorie restricted meals. Their problem is that the meals generate an insulin response. The mice are so insulin sensitive that calories are lost in to adipocytes (and probably hepatocytes) under the over-effective action of insulin.
They lose calories in to adipocytes. These are calories out. The adipocytes get bigger with the lost fat.
Torpor occurs BECAUSE the mice have become fatter.
This is the equivalent of the hunger which follows for a human under a euglycaemic (or even hyperglycaemic) hyperinsulinaemic clamp. There is no hypoglycaemia but fatty acids become locked in to adipocytes by the hyperisulinaemia and hunger follows due a lack of available calories. I posted about it here.
At two hours in to the light period insulin drops low enough to allow lipolysis. The mice wake up.
That's all.
Except: Why do the CR mice have paradoxically (although ns) elevated fasting insulin cf the ad lib mice? There are two reasons. Here are the blood sample times added to the EE graph. The arrows are not quite in the correct clock times, as detailed in the methods, but the times related to feeding/metabolism are approximately correct.
The green arrow is the sampling time for the ad-lib fed mice. It is about six hours in to the light period and the mice would normally have been asleep during the hours leading up to it. Light-period snacking, from the respiratory exchange ratio (RER) graph in Fig3 D, would not normally have started by this time so it's a very simple physiological fasting sample.
The blue arrow for the fasting CR mice just hits the end of torpor. I'm not sure these mice ever have a time when they wouldn't eat, given the chance, but here they are in their hypometabolic phase and have minimal access to calories. At this time insulin is actually a little (ns) higher than for the ad-lib groups (Fig1 D). Higher insulin means fat stays in adipocytes. Why is insulin high?
Calorie restriction does many things in addition to dropping metabolic rate. If you fast a hard working group of humans for 5 days they develop a post prandial increase in GIP (glucose-dependent insulinotropic hormone). This was found in the CR mice in both the fasting and fed state (Fig4 C). GIP facilitates insulin release, hence insulin is a little higher the CR mice and loss of calories in to adipocytes more severe, necessitating torpor.
It's interesting as to why GIP might be elevated under hunger conditions. Possibly generating and saving fat becomes a priority when calories are low. The Ins1+/- CR mice certainly have the highest RER (>1.05) after their third meal, suggesting that they prioritise the conversion of glucose to fat. This DNL, should it occur in the liver, might go some way to explaining the elevated triglycerides in both of the CR groups. Maybe. Accentuated DNL in people who have undergone massive weight loss via gastric bypass surgery is routine during an OGTT. Like these people.
Anyway, I'll stop now. This post is about 1/4 the length it started out as, so if corners seem a bit cut then mea culpa.
Peter
Thursday, September 05, 2019
Hyperlipid Protons ambassadors
Many people may have noticed that the blog Hyperlipid is not exactly the most user friendly of blogs. The prose clearly appears to make sense but some of the concepts are not always particularly simple unless you have the Protons idea well understood.
Last year (2018) Mike Eades made a sterling presentation which summarised the concept in a talk at Low Carb Down Under in terms that were much more accessible
and Brad Marshall now has a blog on which, throughout 2019, he has been writing around ideas which derive partly from the Protons thread on Hyperlipid. But in significantly more user friendly language, while still being on the spot.
Fire in a Bottle
is his website, a very neat name. It's good. He farms low PUFA pigs too.
Peter
*Very few things in life are quite so disappointing as finding that insulin interacts directly with the NOX4 (NADH oxidase 4) complex to generate the bulk of the initiating low levels of superoxide/H2O2 which trigger insulin signalling. Some ROS do come from the electron transport chain but NOX4 seems rather important. Sigh. Bulk ROS to terminate/blunt insulin signalling do appear to be ETC derived...
Last year (2018) Mike Eades made a sterling presentation which summarised the concept in a talk at Low Carb Down Under in terms that were much more accessible
and Brad Marshall now has a blog on which, throughout 2019, he has been writing around ideas which derive partly from the Protons thread on Hyperlipid. But in significantly more user friendly language, while still being on the spot.
Fire in a Bottle
is his website, a very neat name. It's good. He farms low PUFA pigs too.
Peter
*Very few things in life are quite so disappointing as finding that insulin interacts directly with the NOX4 (NADH oxidase 4) complex to generate the bulk of the initiating low levels of superoxide/H2O2 which trigger insulin signalling. Some ROS do come from the electron transport chain but NOX4 seems rather important. Sigh. Bulk ROS to terminate/blunt insulin signalling do appear to be ETC derived...
Sunday, September 01, 2019
The sweet taste of DMEM
I mentioned in the last post that few papers ever specify the glucose concentration used for cell culture. This came up in comments from Alex and I think it deserves a mention in a post of its own.
The standard methods description for almost all cell culture usually specifies that DMEM was used, along with assorted ancillary chemicals and a source of growth factors, usually foetal bovine serum, but rarely the glucose concentration.
A brief trip to any commercial supplier's website gives you approximately 50 DMEM formulations. Three or four have zero glucose and are intended to allow you to specify your own glucose concentration. Another five or six use the original 1g/l of glucose giving 5mmol/l, ie a physiological glucose concentration. This is described as "low glucose".
The other forty-odd specify 4.5g/l, ie 25mmol/l.
I've never done cell culture but I gather the normal technique is to use 25mmol/l medium, let the cells use the glucose to grow and when the glucose drops toward 10mmol/l then they are re-fed with new medium at 25mmol/l of glucose. This works. For decades.
It also provides you with enormous information about the behaviour of cells under hyperglycaemic conditions.
Obviously, 25mmol/l of glucose in an intact human is pure pathology but it is well tolerated by the cells in culture because there is no other caloric source which inputs as FADH2. No free fatty acids.
At the most basic level a glucose concentration of 25mmol/l should a) never occur at all and b) if it does occur it should promptly trigger, via insulin acting on adipocytes, a fall in FFAs to 100 micromol/l or less.
This graph is from a paper on pancreatic beta cell death. From the open circles it is clear that, with glucose held at physiological concentration of 5mmol/l (G5), palmitate is harmless at up to at least 400micromol/l.

This should surprise no one because a 60 hour fast in a healthy human will provide 2000micromol/l of mixed free fatty acids with a glucose of just under 5mmol/l. And no multiple organ failure.
Compare that with the closed circles where glucose is pathologically high at 20mmol/l. Physiological fasting levels of FFAs then unmask the gross pathology of glucose at 20mmol/l. Typical of ordinary cell culture concentration.
Given the idiotic stand of the cardiologist community against saturated fat and given the apparent safety of pathologically elevated glucose in fat-free or PUFA supplemented cell culture medium it should come as no surprise that very few labs worry about the consequences of their grossly non-physiological DMEM.
Demonstrating the pathology of hyperglycaemia using (and blaming) saturated fatty acids or the converse lack of acute toxicity from PUFA (low FADH2 input) is a route to funding which would surely discourage any questioning of the techniques of cell culture which have been successfully used for decades. As folks say:
"We've always done it that way".
That's all I really wanted to say but here are a couple of odds and ends:
There are groups for whom glucose matters. Even if stuck in the incorrect complex I inhibition paradigm for metformin's action, the concentration of glucose is recognised as important by some.
This group:
Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides
have developed a system for glucose homeostasis in cell culture
Their approach is pretty well unique. It is NOT how cell culture is normally done.
The other brief aside while I'm on cell culture from an outsider viewpoint is FBS, foetal bovine serum. This contains essential growth factors. You can pretty well translate it as "insulin" or "IGF-1".
When cell culture shows that metformin works to reduce cancer cell growth, it is working in the presence of insulin signalling (usually combined with hyperglycaemia). Which metformin blocks at the glycerophosphate shuttle level, without the need for a lethal blockade of complex I.
Peter
The standard methods description for almost all cell culture usually specifies that DMEM was used, along with assorted ancillary chemicals and a source of growth factors, usually foetal bovine serum, but rarely the glucose concentration.
A brief trip to any commercial supplier's website gives you approximately 50 DMEM formulations. Three or four have zero glucose and are intended to allow you to specify your own glucose concentration. Another five or six use the original 1g/l of glucose giving 5mmol/l, ie a physiological glucose concentration. This is described as "low glucose".
The other forty-odd specify 4.5g/l, ie 25mmol/l.
I've never done cell culture but I gather the normal technique is to use 25mmol/l medium, let the cells use the glucose to grow and when the glucose drops toward 10mmol/l then they are re-fed with new medium at 25mmol/l of glucose. This works. For decades.
It also provides you with enormous information about the behaviour of cells under hyperglycaemic conditions.
Obviously, 25mmol/l of glucose in an intact human is pure pathology but it is well tolerated by the cells in culture because there is no other caloric source which inputs as FADH2. No free fatty acids.
At the most basic level a glucose concentration of 25mmol/l should a) never occur at all and b) if it does occur it should promptly trigger, via insulin acting on adipocytes, a fall in FFAs to 100 micromol/l or less.
This graph is from a paper on pancreatic beta cell death. From the open circles it is clear that, with glucose held at physiological concentration of 5mmol/l (G5), palmitate is harmless at up to at least 400micromol/l.

This should surprise no one because a 60 hour fast in a healthy human will provide 2000micromol/l of mixed free fatty acids with a glucose of just under 5mmol/l. And no multiple organ failure.
Compare that with the closed circles where glucose is pathologically high at 20mmol/l. Physiological fasting levels of FFAs then unmask the gross pathology of glucose at 20mmol/l. Typical of ordinary cell culture concentration.
Given the idiotic stand of the cardiologist community against saturated fat and given the apparent safety of pathologically elevated glucose in fat-free or PUFA supplemented cell culture medium it should come as no surprise that very few labs worry about the consequences of their grossly non-physiological DMEM.
Demonstrating the pathology of hyperglycaemia using (and blaming) saturated fatty acids or the converse lack of acute toxicity from PUFA (low FADH2 input) is a route to funding which would surely discourage any questioning of the techniques of cell culture which have been successfully used for decades. As folks say:
"We've always done it that way".
That's all I really wanted to say but here are a couple of odds and ends:
There are groups for whom glucose matters. Even if stuck in the incorrect complex I inhibition paradigm for metformin's action, the concentration of glucose is recognised as important by some.
This group:
Metabolic determinants of cancer cell sensitivity to glucose limitation and biguanides
have developed a system for glucose homeostasis in cell culture
Their approach is pretty well unique. It is NOT how cell culture is normally done.
The other brief aside while I'm on cell culture from an outsider viewpoint is FBS, foetal bovine serum. This contains essential growth factors. You can pretty well translate it as "insulin" or "IGF-1".
When cell culture shows that metformin works to reduce cancer cell growth, it is working in the presence of insulin signalling (usually combined with hyperglycaemia). Which metformin blocks at the glycerophosphate shuttle level, without the need for a lethal blockade of complex I.
Peter
Wednesday, August 28, 2019
Who gave you pancreatitis?
Background: Propofol is a mainstay anaesthetic induction agent. Its use is associated with occasional pancreatitis episodes. I won't wander aside about the poor dog which was referred for cataract surgery and which did eventually recovered from its perioperative fulminating pancreatitis.
Propofol is dissolved in what is essentially Intralipid, a soybean emulsion used for parenteral nutrition. The lipid emulsion gives a transient hypertriglyceridaemia. Hypertriglyceridaemia from any cause is associated with pancreatitis.
Me, being me, would automatically blame the PUFA in the Intralipid. But then I would.
I came across this paper by accident this morning:
Distinctive roles of unsaturated and saturated fatty acids in hyperlipidemic pancreatitis
It's good. The group even give you the glucose concentration used in their cell culture, 0.2%, ie 200mg/dl or around 10mmol/l. Not normal but hardly seriously pathological.
Aside: Less innocent groups keep quiet about glucose concentration and can reliably show endoplasmic reticulum stress and any other nasty attributable to palmitic acid. With how much glucose??? End aside.
This is what they found:
"Unsaturated fatty acids at high concentrations but not saturated fatty acids induced intra-acinar cell trypsin activation and cell damage and increased PKC expression"
So. If you have a genetic hypertriglyceridaemia, say lipoprotein lipase deficiency, and you get acute necrotising pancreatitis (not fun) there is every possibility that it was induced by the high content of polyunsaturated fatty acids in those triglycerides and the FFAs derived from them.
Which means that your cardiologist put you in the ITU. Avoid saturated fats, replace them with polyunsaturated oils. Thank you Public Health England and your equivalents world wide.
The converse might well be that loss of the gene for lipoprotein lipase, or similar loss, might not have been a big deal when humans lived by eating elephants. Or even until corn oil took off as a cholesterol lowering scam.
Peter
Propofol is dissolved in what is essentially Intralipid, a soybean emulsion used for parenteral nutrition. The lipid emulsion gives a transient hypertriglyceridaemia. Hypertriglyceridaemia from any cause is associated with pancreatitis.
Me, being me, would automatically blame the PUFA in the Intralipid. But then I would.
I came across this paper by accident this morning:
Distinctive roles of unsaturated and saturated fatty acids in hyperlipidemic pancreatitis
It's good. The group even give you the glucose concentration used in their cell culture, 0.2%, ie 200mg/dl or around 10mmol/l. Not normal but hardly seriously pathological.
Aside: Less innocent groups keep quiet about glucose concentration and can reliably show endoplasmic reticulum stress and any other nasty attributable to palmitic acid. With how much glucose??? End aside.
This is what they found:
"Unsaturated fatty acids at high concentrations but not saturated fatty acids induced intra-acinar cell trypsin activation and cell damage and increased PKC expression"
So. If you have a genetic hypertriglyceridaemia, say lipoprotein lipase deficiency, and you get acute necrotising pancreatitis (not fun) there is every possibility that it was induced by the high content of polyunsaturated fatty acids in those triglycerides and the FFAs derived from them.
Which means that your cardiologist put you in the ITU. Avoid saturated fats, replace them with polyunsaturated oils. Thank you Public Health England and your equivalents world wide.
The converse might well be that loss of the gene for lipoprotein lipase, or similar loss, might not have been a big deal when humans lived by eating elephants. Or even until corn oil took off as a cholesterol lowering scam.
Peter
Protons (50) The video
Dave Speijer has a great video up on YouTube based largely around his recent paper in BioEssays.
Can All Major ROS Forming Sites of the Respiratory Chain Be Activated By High FADH2/NADH Ratios?
It's nice to hear the man himself on a subject about which he has spent a great deal of time thinking. Here it is:
Many thanks to Andrew Moore, editor-in-chief at BioEssays and publisher of many of Dr Speijer's papers, for the heads up that this video had been produced.
Peter
Can All Major ROS Forming Sites of the Respiratory Chain Be Activated By High FADH2/NADH Ratios?
It's nice to hear the man himself on a subject about which he has spent a great deal of time thinking. Here it is:
Many thanks to Andrew Moore, editor-in-chief at BioEssays and publisher of many of Dr Speijer's papers, for the heads up that this video had been produced.
Peter
Monday, August 19, 2019
Protons (49) Complex III
Dave Speijer, an extremely insightful person if ever there was one, has a new paper out:
Can All Major ROS Forming Sites of the Respiratory Chain Be Activated By High FADH2/NADH Ratios?
the link to which I am extremely grateful to Bob for forwarding to me. This concept is purely from Dr Speijer. But I like it. A lot.
I'll start with an old doodle I produced about a decade ago depicting the front end of the electron transport chain. Matrix is at the top, cytoplasm at the bottom:
Electrons travel from NADH to Coenzyme Q, reducing it to QH2 which donates them to complex III being oxidised back to Q in the process.
The unlabelled blobs in the diagram are complex II (succinate dehydrogenase), electron transporting flavoprotein dehydrogenase and mitochondrial glycerophosphate dehydrogenase, all of which compete with complex I for CoQ as an electron acceptor. Given a high membrane voltage, a deeply reduced CoQ couple with most of the CoQ as QH2, then reverse electron transport back through complex I gives superoxide/H2O2 generation (provided the mitochondrial NAD pool is also highly reduced so unable to accept electrons (ie little NAD+, lots of NADH).
This is pretty straight forward and is the gist of the Protons thread. The extension of this is that, under high substrate availability, H2O2 from this process stops insulin facilitated caloric ingress to the cell.
Another major site of ROS generation in the ETC is complex III. Dave Speijer would hold that this is also triggered, like RET, by a deeply reduced CoQ couple. This is why.
So here is a stripped out version of the above doodle:
The problem here is that it's not that simple. Complex III does rather odd things with its two electrons. Electron bifurcation is a standard enzymic technique perfected very early on by proto-biology and it is exactly what happens here. One electron travels to cytochrome C (which only ever carries one electron at a time) and then on to complex IV and O2. The other does not:
The second electron is transferred backwards to another CoQ molecule and so partially reduces it to QH*, the radical semiquinone.
So (oxidised) Q is a necessary electron acceptor for complex III. Under high substrate availability and with most of the CoQ couple present as QH2 there will be very little Q available.
With one electron securely on cytochrome C, with any delay in the availability of Q, the second electron can be left sitting on one of the two haem groups along its route, easily available for donation to O2 to give superoxide, so adding its ROS signal to that of complex I. Both indicate that enough substrate is present and it is time to limit insulin signalling, to limit caloric ingress.
Nothing, absolutely nothing, about the construction of the ETC is random.
The general principle that a highly reduced CoQ couple is a signal to halt caloric ingress in to the cell applies to complex III just as much as to complex I.
When you want to resist insulin, you really want to resist insulin. Superoxide is then your friend.
If you fail to limit caloric ingress then eventually ROS from complex III are ideally placed to break the cardiolipins which anchor the water soluble cytochrome C to the inner mitochondrial membrane. Destroy these anchors and the ROS signal changes from"resist insulin" (good) to "perform apoptosis" (possibly not quite so good)...
And of course ROS in excess of physiological signalling are going to activate all sorts of inflammatory pathways.
Peter
Aside: The process of ROS generation is probably limited by pairing of complex IIIs (complex III is always a dimer in-vivo) for cooperation to use one Q to accept an electron from each of the pair. This should limit ROS generation as one Q will be available to two complex IIIs, twice that avaiable if they were each working alone. Nothing is random. End aside.
Can All Major ROS Forming Sites of the Respiratory Chain Be Activated By High FADH2/NADH Ratios?
the link to which I am extremely grateful to Bob for forwarding to me. This concept is purely from Dr Speijer. But I like it. A lot.
I'll start with an old doodle I produced about a decade ago depicting the front end of the electron transport chain. Matrix is at the top, cytoplasm at the bottom:
Electrons travel from NADH to Coenzyme Q, reducing it to QH2 which donates them to complex III being oxidised back to Q in the process.
The unlabelled blobs in the diagram are complex II (succinate dehydrogenase), electron transporting flavoprotein dehydrogenase and mitochondrial glycerophosphate dehydrogenase, all of which compete with complex I for CoQ as an electron acceptor. Given a high membrane voltage, a deeply reduced CoQ couple with most of the CoQ as QH2, then reverse electron transport back through complex I gives superoxide/H2O2 generation (provided the mitochondrial NAD pool is also highly reduced so unable to accept electrons (ie little NAD+, lots of NADH).
This is pretty straight forward and is the gist of the Protons thread. The extension of this is that, under high substrate availability, H2O2 from this process stops insulin facilitated caloric ingress to the cell.
Another major site of ROS generation in the ETC is complex III. Dave Speijer would hold that this is also triggered, like RET, by a deeply reduced CoQ couple. This is why.
So here is a stripped out version of the above doodle:
The problem here is that it's not that simple. Complex III does rather odd things with its two electrons. Electron bifurcation is a standard enzymic technique perfected very early on by proto-biology and it is exactly what happens here. One electron travels to cytochrome C (which only ever carries one electron at a time) and then on to complex IV and O2. The other does not:
The second electron is transferred backwards to another CoQ molecule and so partially reduces it to QH*, the radical semiquinone.
So (oxidised) Q is a necessary electron acceptor for complex III. Under high substrate availability and with most of the CoQ couple present as QH2 there will be very little Q available.
With one electron securely on cytochrome C, with any delay in the availability of Q, the second electron can be left sitting on one of the two haem groups along its route, easily available for donation to O2 to give superoxide, so adding its ROS signal to that of complex I. Both indicate that enough substrate is present and it is time to limit insulin signalling, to limit caloric ingress.
Nothing, absolutely nothing, about the construction of the ETC is random.
The general principle that a highly reduced CoQ couple is a signal to halt caloric ingress in to the cell applies to complex III just as much as to complex I.
When you want to resist insulin, you really want to resist insulin. Superoxide is then your friend.
If you fail to limit caloric ingress then eventually ROS from complex III are ideally placed to break the cardiolipins which anchor the water soluble cytochrome C to the inner mitochondrial membrane. Destroy these anchors and the ROS signal changes from"resist insulin" (good) to "perform apoptosis" (possibly not quite so good)...
And of course ROS in excess of physiological signalling are going to activate all sorts of inflammatory pathways.
Peter
Aside: The process of ROS generation is probably limited by pairing of complex IIIs (complex III is always a dimer in-vivo) for cooperation to use one Q to accept an electron from each of the pair. This should limit ROS generation as one Q will be available to two complex IIIs, twice that avaiable if they were each working alone. Nothing is random. End aside.
Subscribe to:
Posts (Atom)